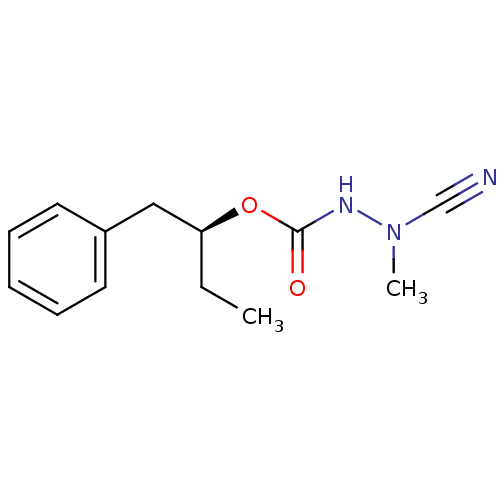
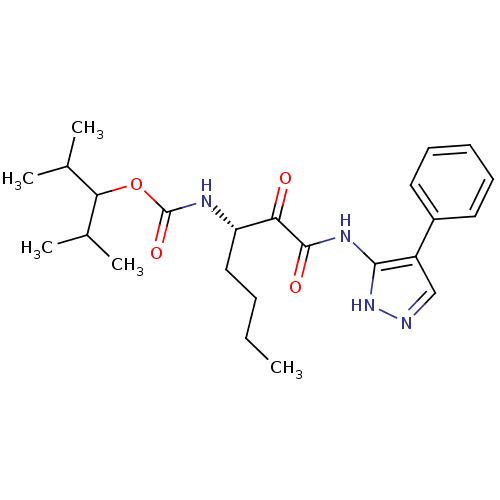
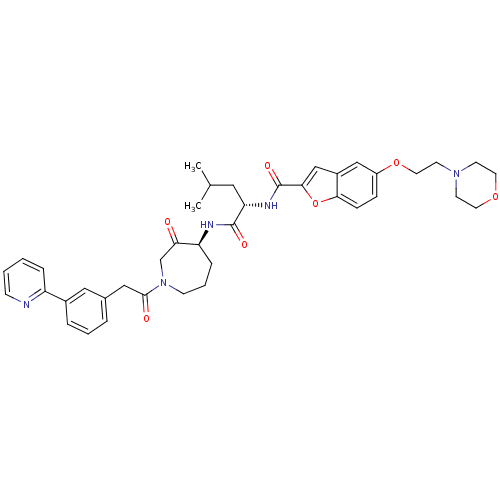
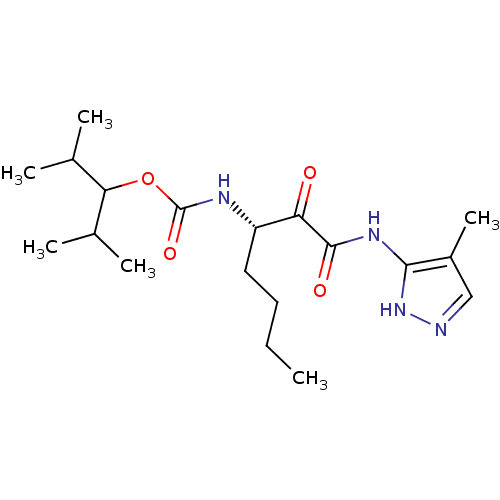
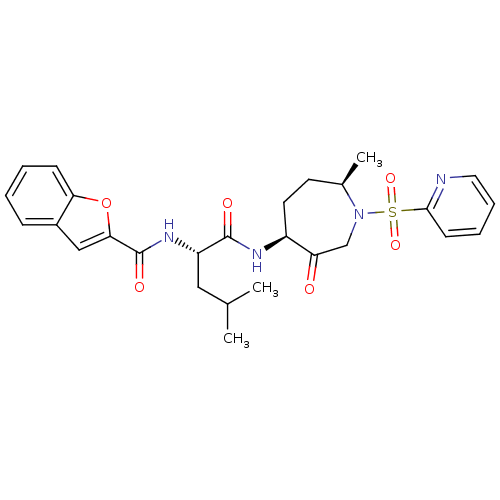
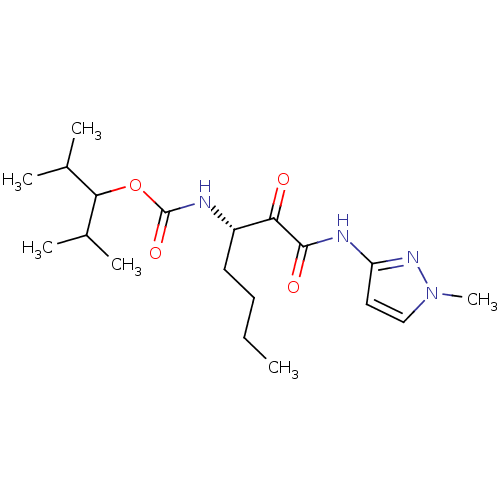
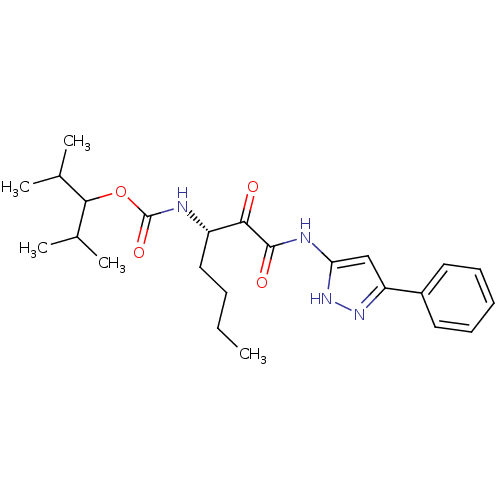
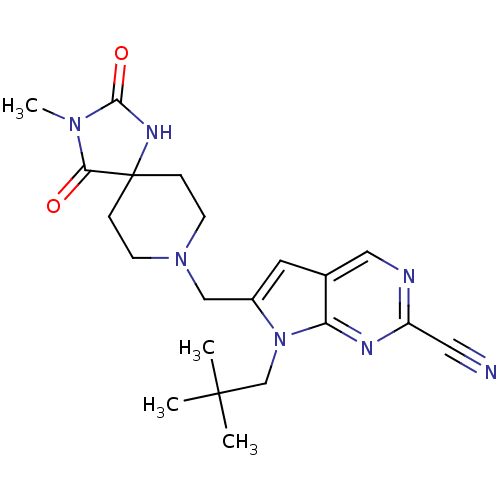
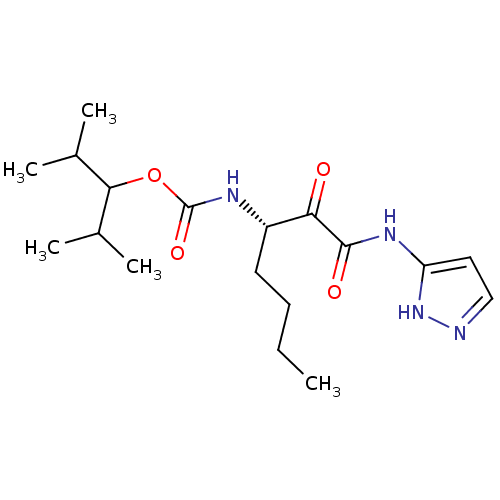
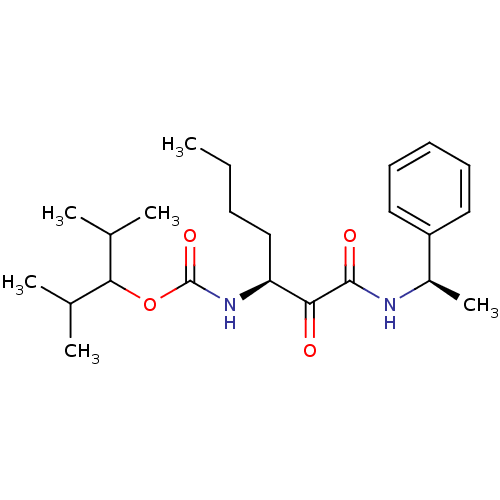
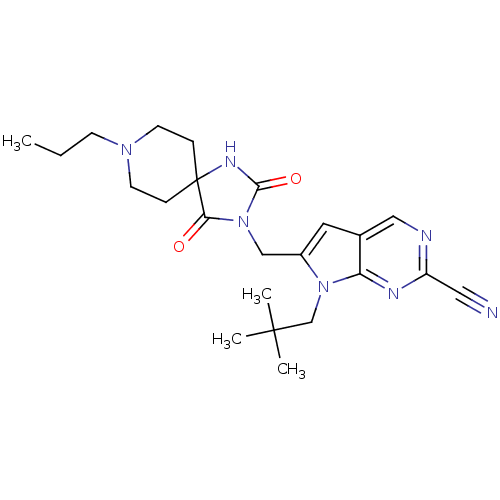
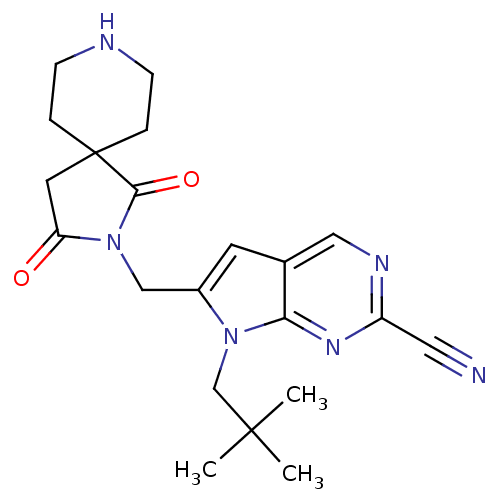
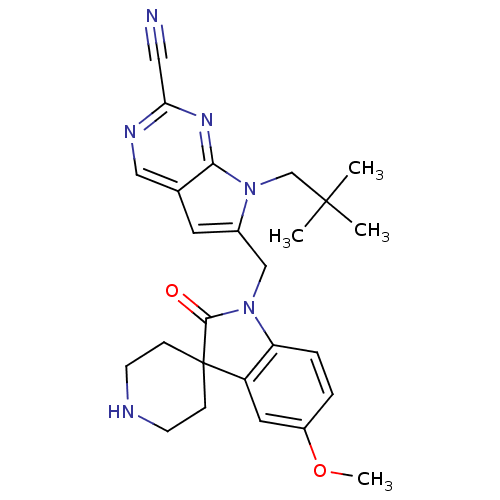
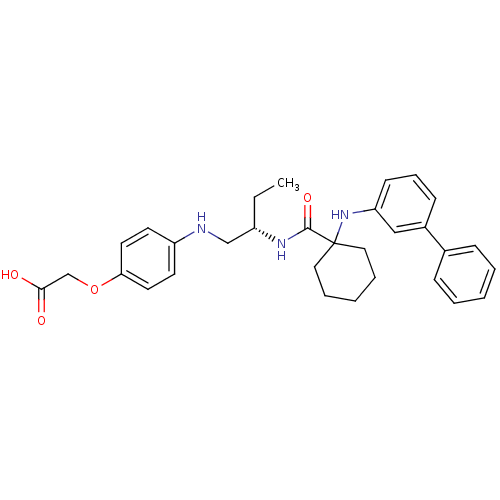
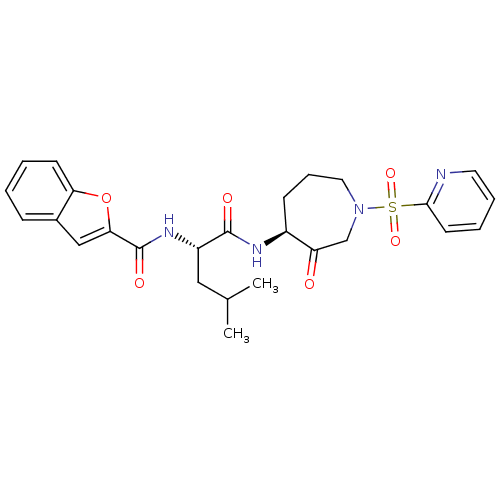
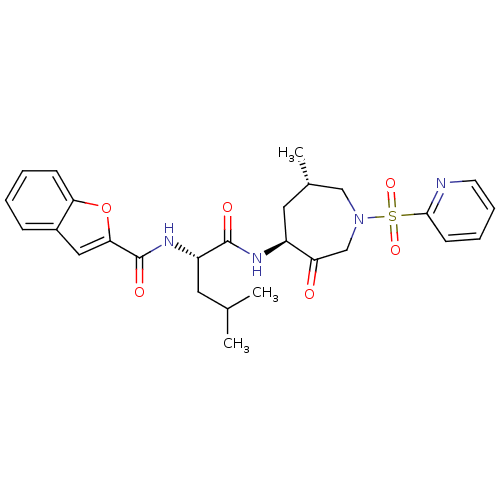
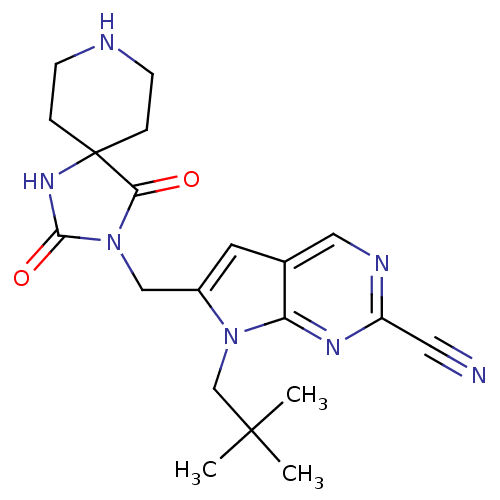
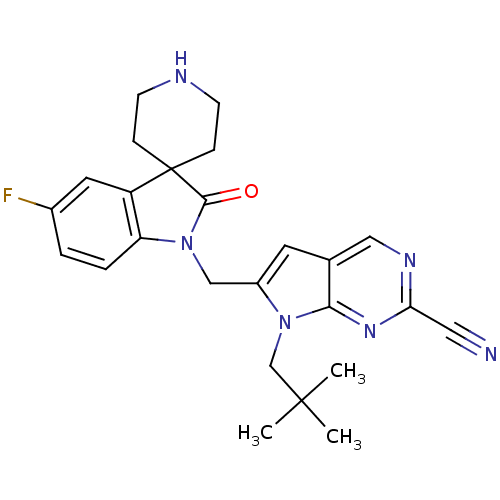
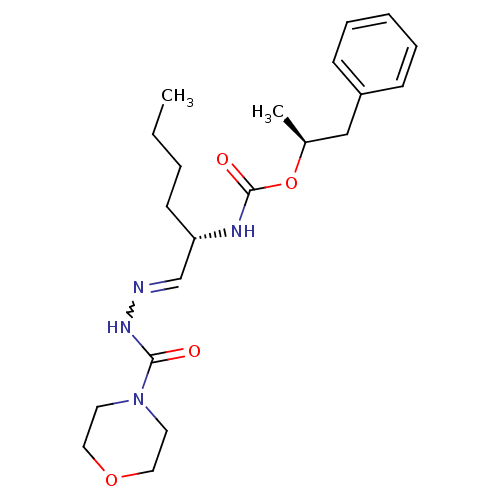
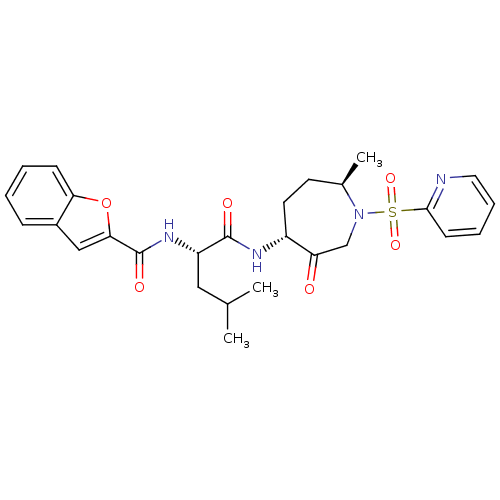
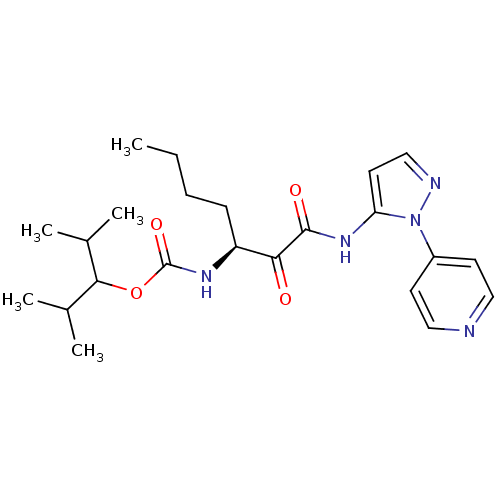
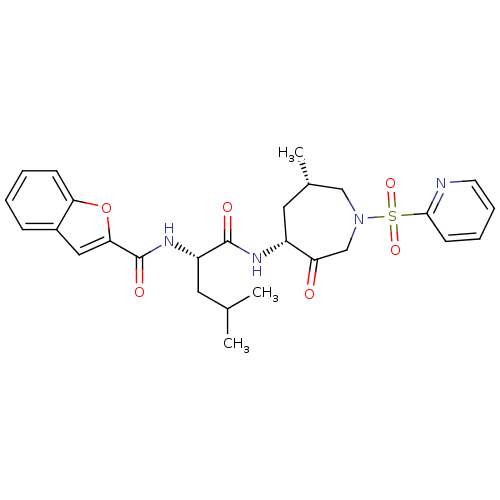
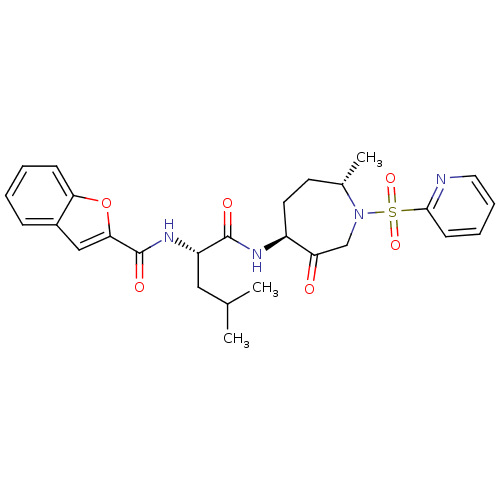
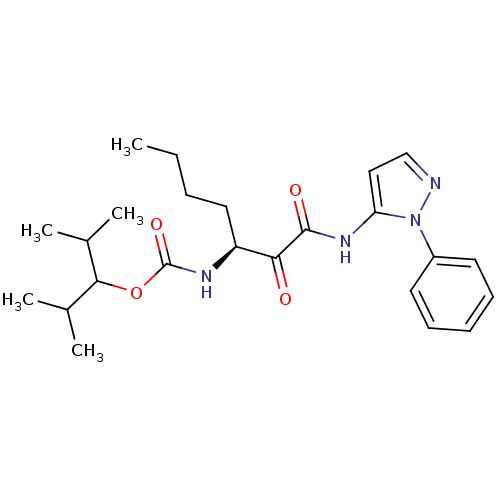
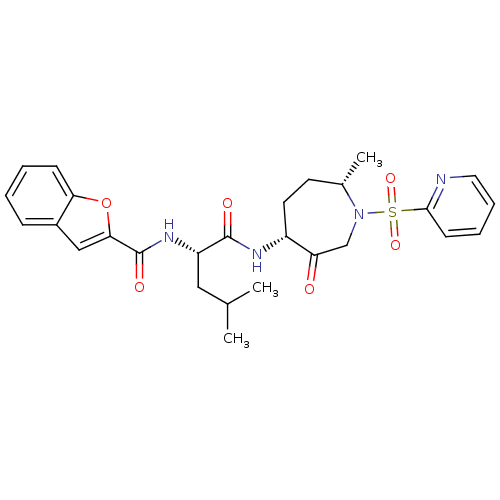
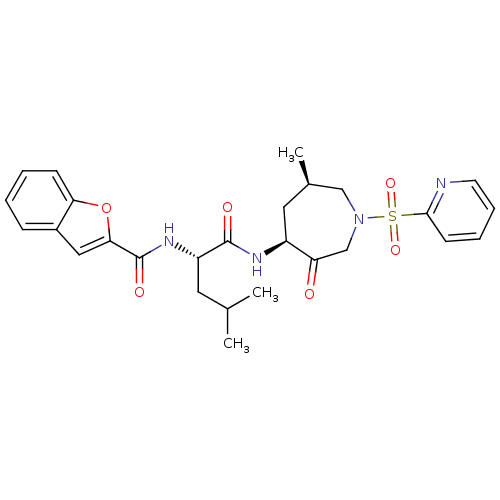
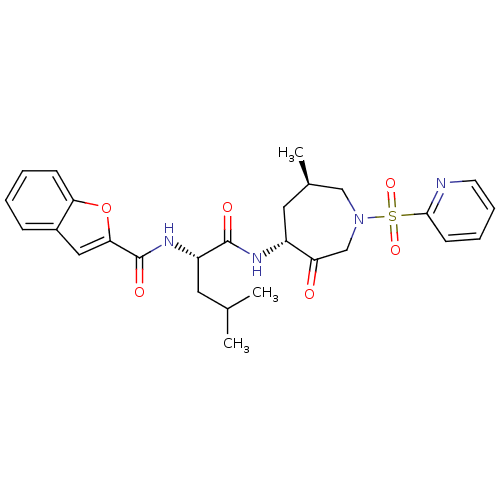
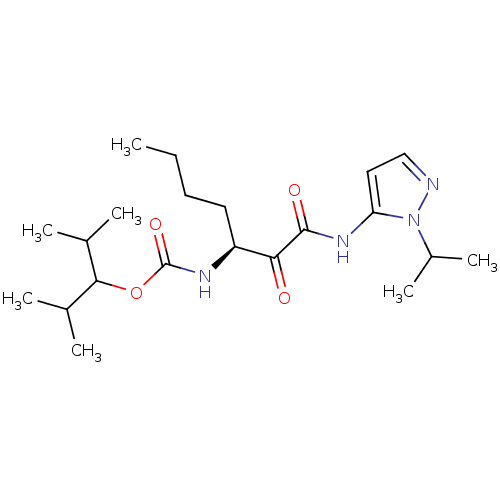
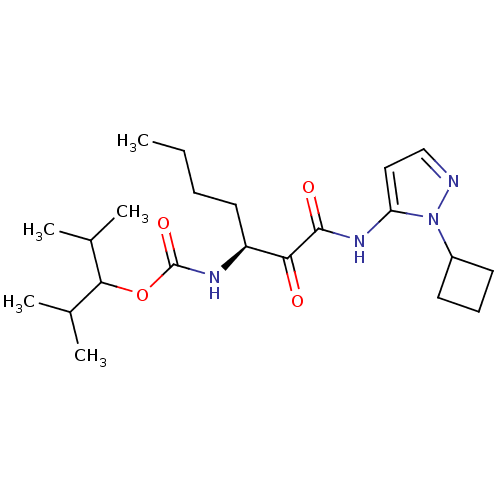
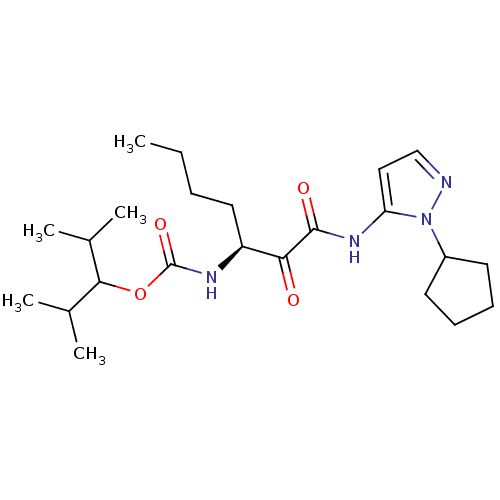
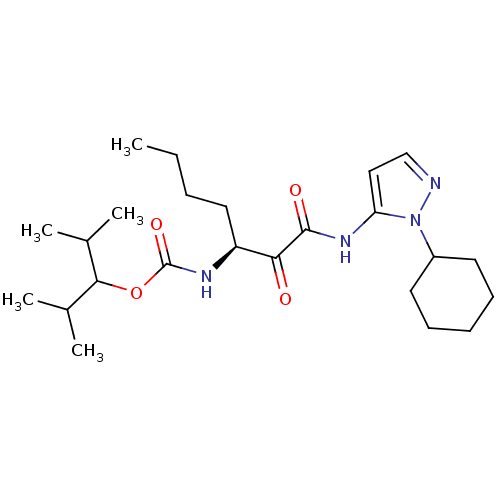
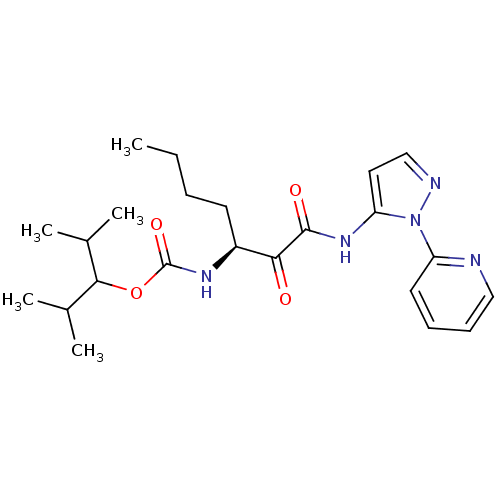
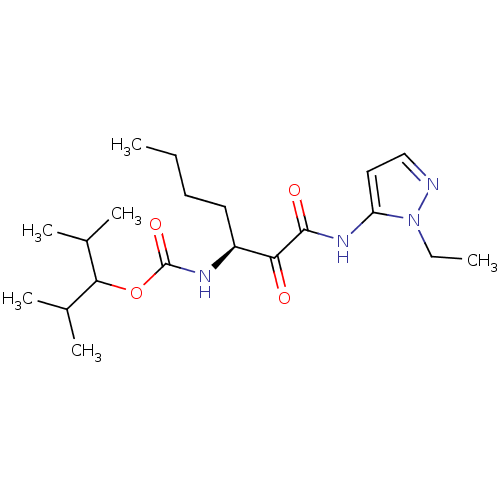
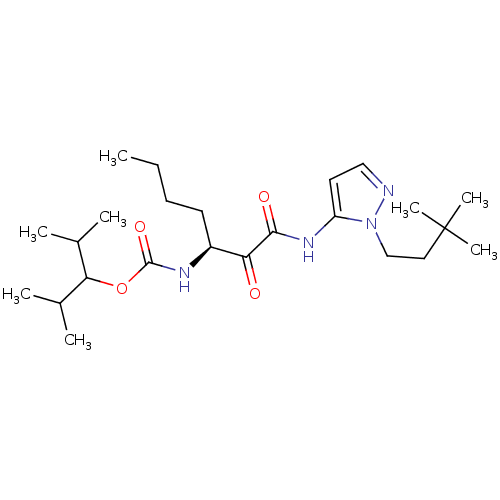
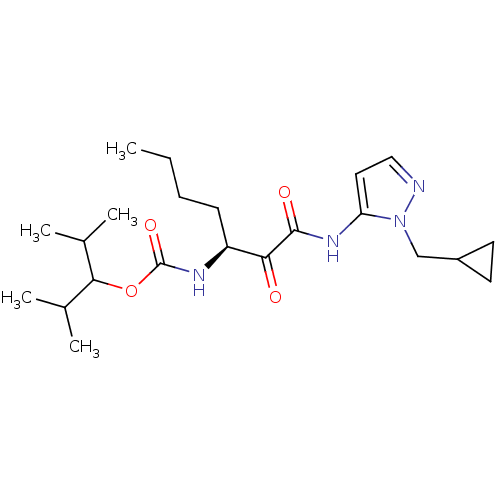
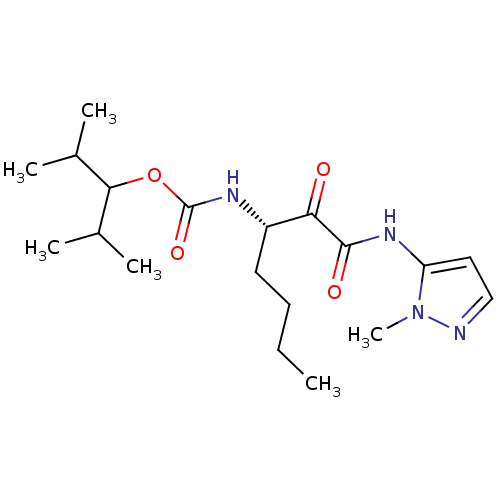
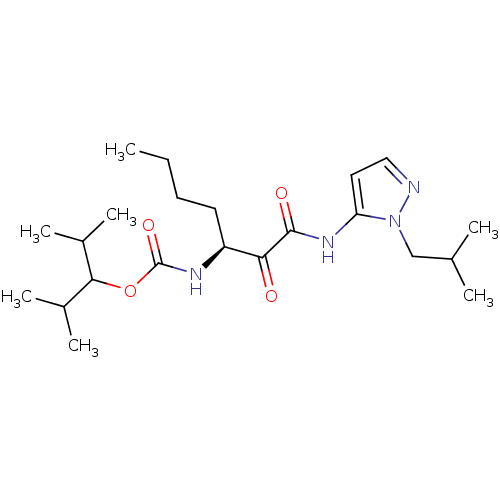
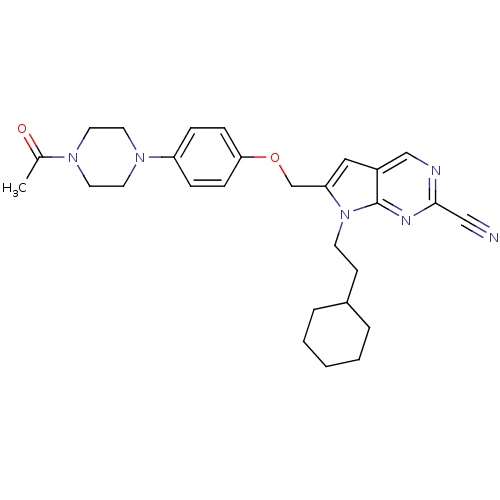
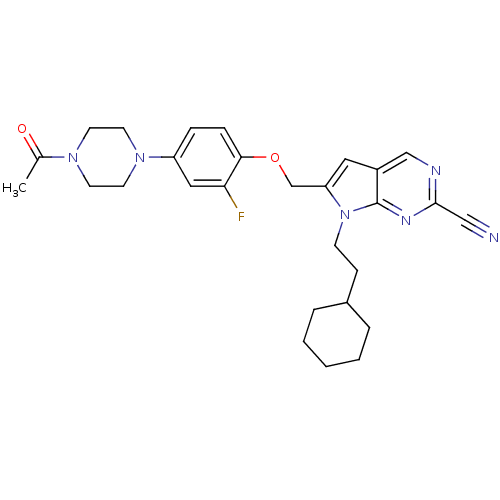
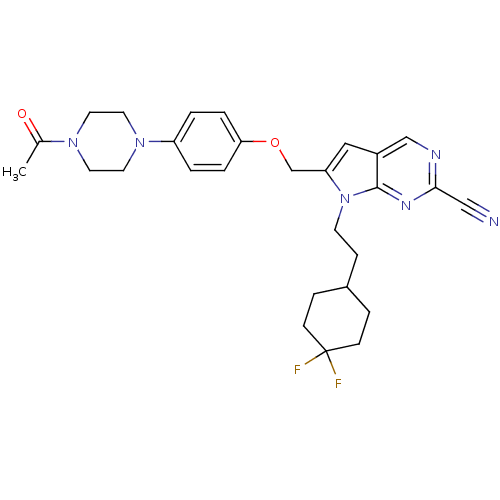
Affinity DataKi: 0.0720nMAssay Description:Inhibition constant against rat cathepsin KMore data for this Ligand-Target Pair
Affinity DataIC50: 1.80nMAssay Description:Inhibition of cysteine protease cathepsin K of ratMore data for this Ligand-Target Pair
Affinity DataKi: 4.80nMAssay Description:Inhibitory activity against Rat cathepsin KMore data for this Ligand-Target Pair
Affinity DataIC50: 5.5nMAssay Description:Inhibition of cysteine protease cathepsin K of ratMore data for this Ligand-Target Pair
Affinity DataKi: 8nMAssay Description:Potential inhibitors were evaluated using the progress curve method. Assays were carried out in the presence of variable concentrations of test compo...More data for this Ligand-Target Pair
Affinity DataIC50: 11nMAssay Description:Inhibition of cysteine protease cathepsin K of ratMore data for this Ligand-Target Pair
Affinity DataIC50: 12nMAssay Description:Inhibition of cysteine protease cathepsin K of ratMore data for this Ligand-Target Pair
Affinity DataIC50: 12nMAssay Description:Inhibition of rat recombinant cathepsin K expressed in Sf21 cells by fluorescence assayMore data for this Ligand-Target Pair
Affinity DataIC50: 13nMAssay Description:Inhibition of cysteine protease cathepsin K of ratMore data for this Ligand-Target Pair
Affinity DataIC50: 20nMAssay Description:Inhibitory activity against rat cathepsin KMore data for this Ligand-Target Pair
Affinity DataIC50: 27nMAssay Description:Inhibition of rat recombinant cathepsin K expressed in Sf21 cells by fluorescence assayMore data for this Ligand-Target Pair
Affinity DataIC50: 30nMAssay Description:Inhibition of rat recombinant cathepsin K expressed in Sf21 cells by fluorescence assayMore data for this Ligand-Target Pair
Affinity DataIC50: 44nMAssay Description:Inhibition of rat recombinant cathepsin K expressed in Sf21 cells by fluorescence assayMore data for this Ligand-Target Pair
Affinity DataIC50: 58nMAssay Description:Inhibition of rat cathepsin KMore data for this Ligand-Target Pair
Affinity DataKi: 60nMAssay Description:Potential inhibitors were evaluated using the progress curve method. Assays were carried out in the presence of variable concentrations of test compo...More data for this Ligand-Target Pair
Affinity DataKi: 60nMAssay Description:Inhibitory activity against Rat cathepsin KMore data for this Ligand-Target Pair
Affinity DataKi: 73nMAssay Description:Potential inhibitors were evaluated using the progress curve method. Assays were carried out in the presence of variable concentrations of test compo...More data for this Ligand-Target Pair
Affinity DataIC50: 76nMAssay Description:Inhibition of rat recombinant cathepsin K expressed in Sf21 cells by fluorescence assayMore data for this Ligand-Target Pair
Affinity DataIC50: 78nMAssay Description:Inhibition of rat recombinant cathepsin K expressed in Sf21 cells by fluorescence assayMore data for this Ligand-Target Pair
Affinity DataIC50: 130nMAssay Description:Inhibition of rat cathepsin KMore data for this Ligand-Target Pair
Affinity DataKi: 430nMAssay Description:Potential inhibitors were evaluated using the progress curve method. Assays were carried out in the presence of variable concentrations of test compo...More data for this Ligand-Target Pair
Affinity DataIC50: 470nMAssay Description:Inhibition of cysteine protease cathepsin K of ratMore data for this Ligand-Target Pair
Affinity DataKi: 660nMAssay Description:Potential inhibitors were evaluated using the progress curve method. Assays were carried out in the presence of variable concentrations of test compo...More data for this Ligand-Target Pair
Affinity DataKi: 680nMAssay Description:Potential inhibitors were evaluated using the progress curve method. Assays were carried out in the presence of variable concentrations of test compo...More data for this Ligand-Target Pair
Affinity DataIC50: 790nMAssay Description:Inhibition of cysteine protease cathepsin K of ratMore data for this Ligand-Target Pair
Affinity DataKi: 790nMAssay Description:Potential inhibitors were evaluated using the progress curve method. Assays were carried out in the presence of variable concentrations of test compo...More data for this Ligand-Target Pair
Affinity DataKi: >1.00E+3nMAssay Description:Potential inhibitors were evaluated using the progress curve method. Assays were carried out in the presence of variable concentrations of test compo...More data for this Ligand-Target Pair
Affinity DataKi: >1.00E+3nMAssay Description:Potential inhibitors were evaluated using the progress curve method. Assays were carried out in the presence of variable concentrations of test compo...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+3nMAssay Description:Inhibition of cysteine protease cathepsin K of ratMore data for this Ligand-Target Pair
Affinity DataIC50: 1.10E+3nMAssay Description:Inhibition of cysteine protease cathepsin K of ratMore data for this Ligand-Target Pair
Affinity DataIC50: 1.10E+3nMAssay Description:Inhibition of cysteine protease cathepsin K of ratMore data for this Ligand-Target Pair
Affinity DataIC50: 1.20E+3nMAssay Description:Inhibition of cysteine protease cathepsin K of ratMore data for this Ligand-Target Pair
Affinity DataIC50: 1.20E+3nMAssay Description:Inhibition of cysteine protease cathepsin K of ratMore data for this Ligand-Target Pair
Affinity DataIC50: 1.50E+3nMAssay Description:Inhibition of cysteine protease cathepsin K of ratMore data for this Ligand-Target Pair
Affinity DataIC50: 1.60E+3nMAssay Description:Inhibition of cysteine protease cathepsin K of ratMore data for this Ligand-Target Pair
Affinity DataIC50: 1.70E+3nMAssay Description:Inhibition of cysteine protease cathepsin K of ratMore data for this Ligand-Target Pair
Affinity DataIC50: 1.70E+3nMAssay Description:Inhibition of cysteine protease cathepsin K of ratMore data for this Ligand-Target Pair
Affinity DataIC50: 2.70E+3nMAssay Description:Inhibition of cysteine protease cathepsin K of ratMore data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+4nMpH: 7.0 T: 2°CAssay Description:The substrate hydrolysis with or without inhibitor was monitored at an excitation wavelength of 360nm and an emission wavelength of 460 nm on a fluor...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+4nMpH: 7.0 T: 2°CAssay Description:The substrate hydrolysis with or without inhibitor was monitored at an excitation wavelength of 360nm and an emission wavelength of 460 nm on a fluor...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+4nMpH: 7.0 T: 2°CAssay Description:The substrate hydrolysis with or without inhibitor was monitored at an excitation wavelength of 360nm and an emission wavelength of 460 nm on a fluor...More data for this Ligand-Target Pair