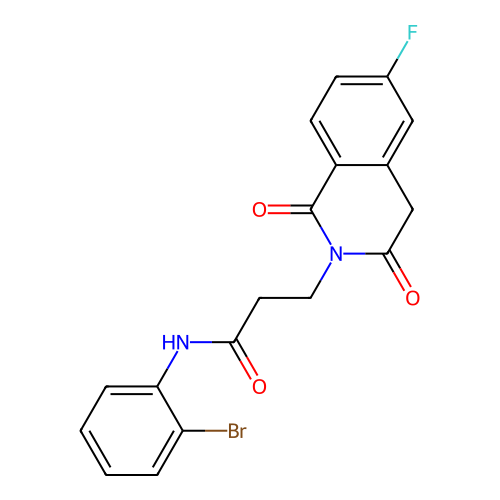
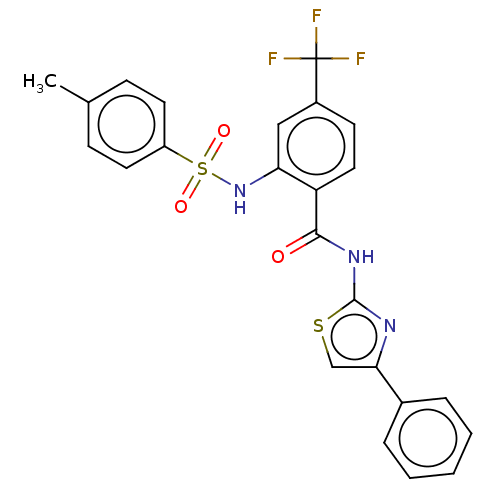
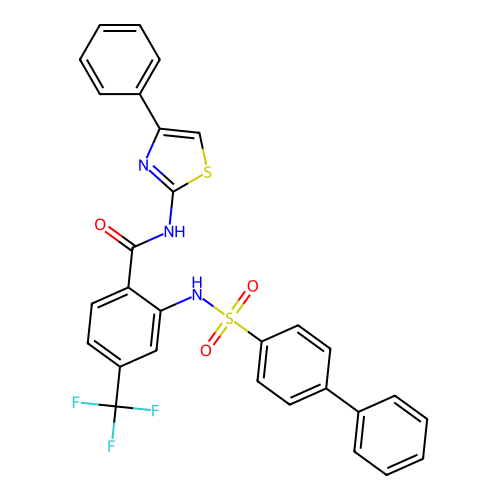
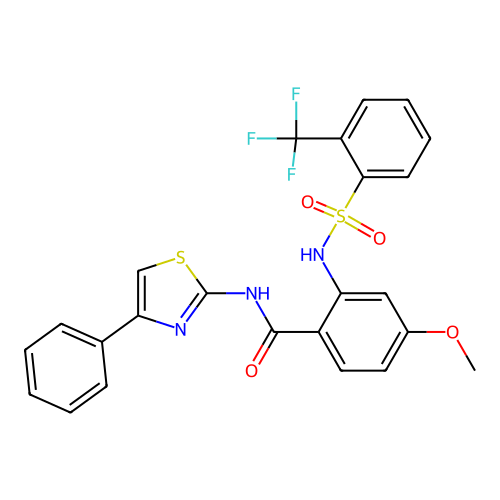
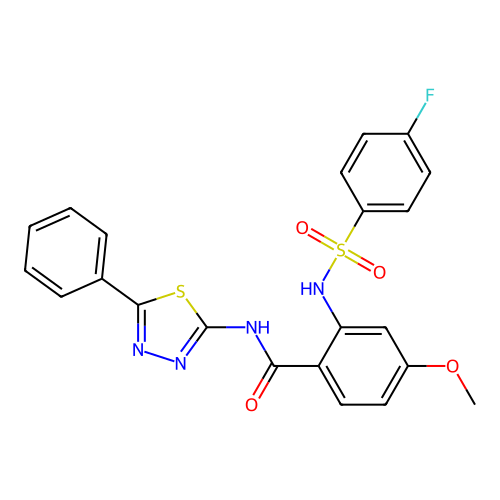
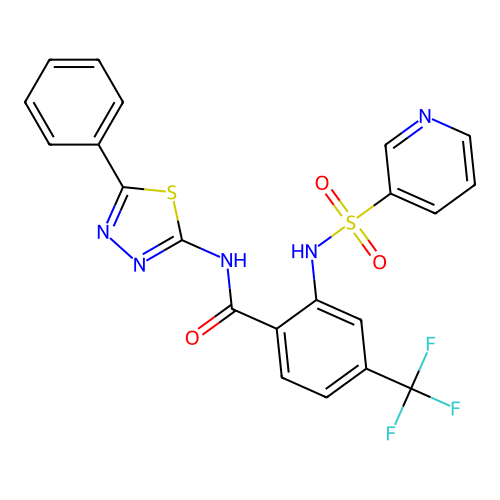
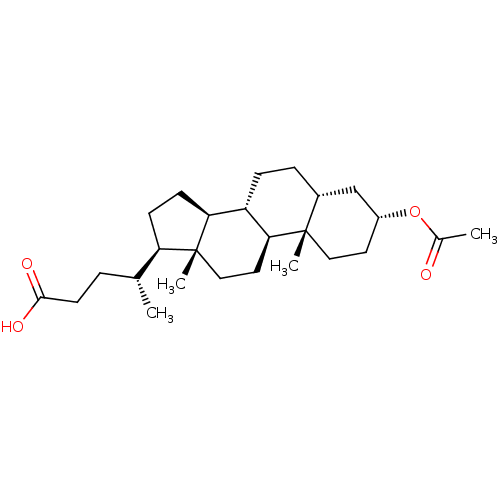
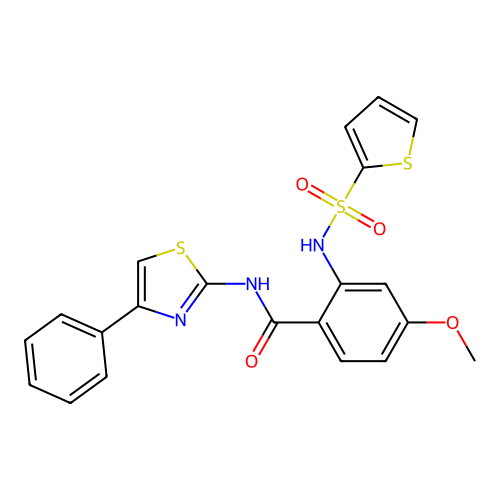
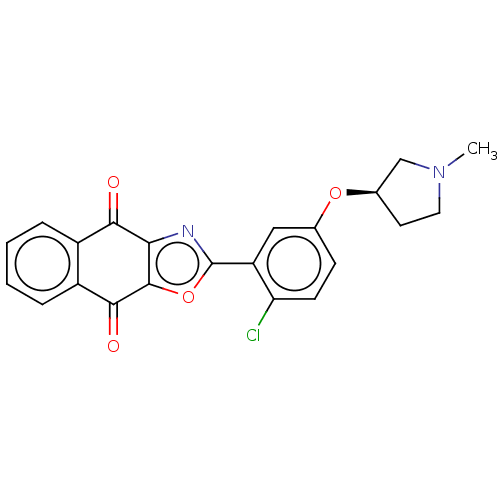
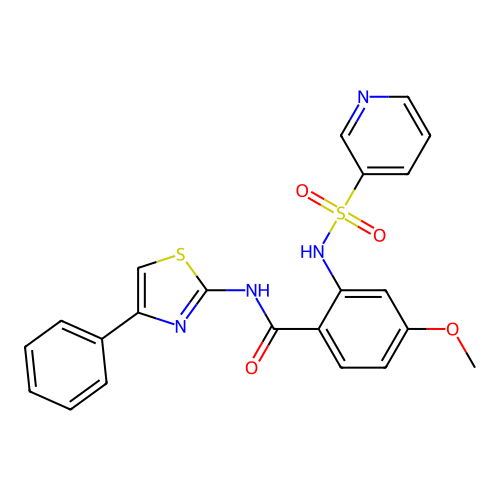
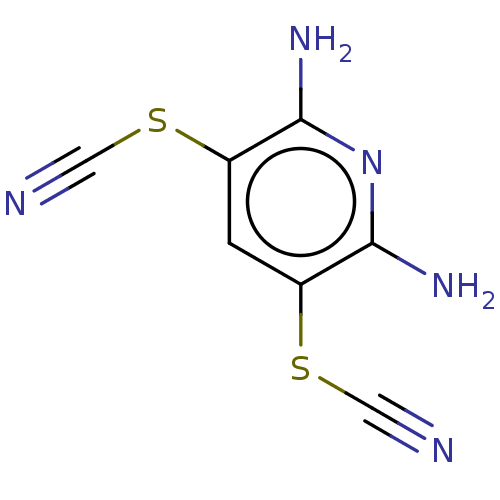
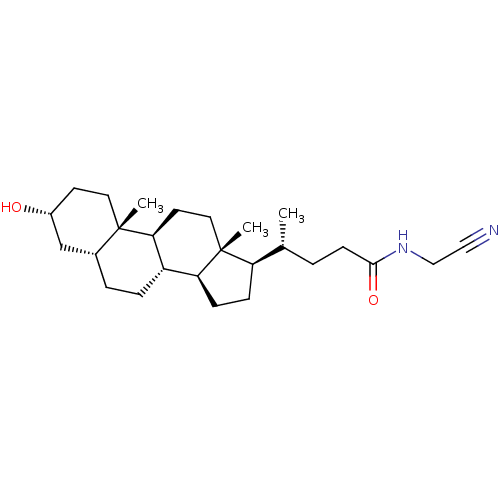
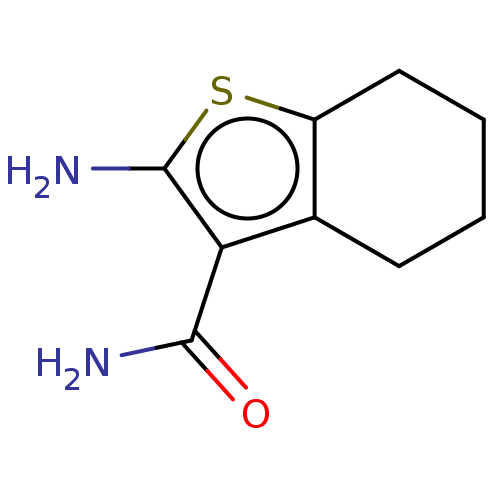
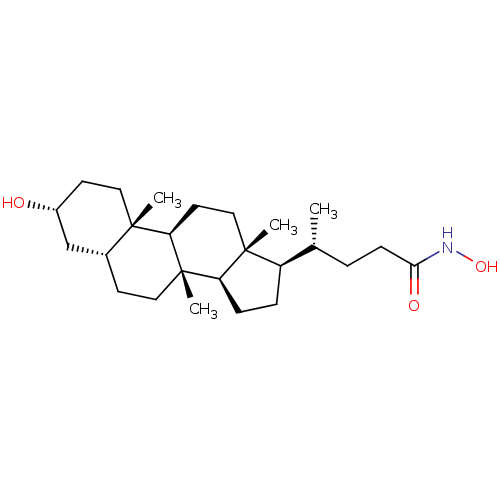
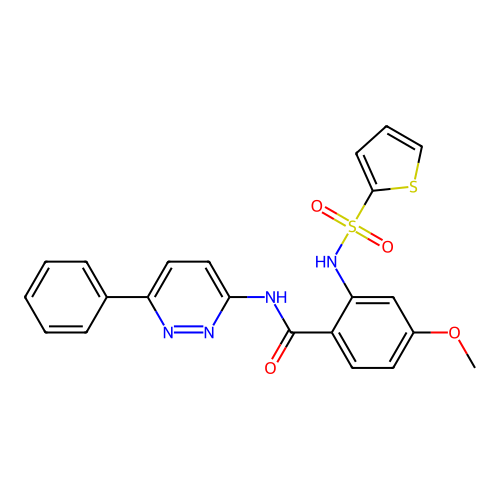
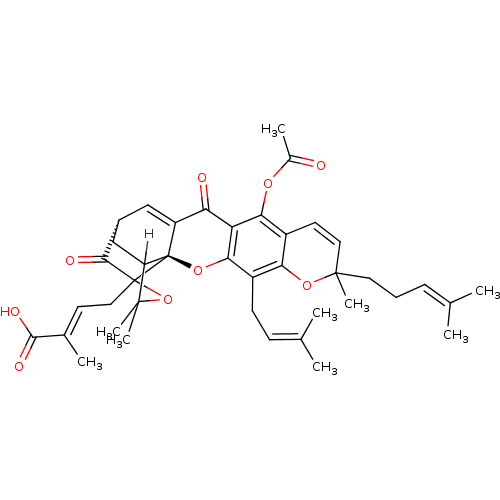
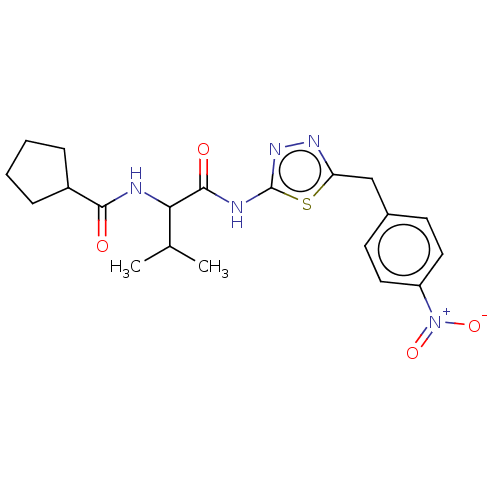
TargetUbiquitin carboxyl-terminal hydrolase 2(Human)
China Pharmaceutical University
Curated by ChEMBL
China Pharmaceutical University
Curated by ChEMBL
Affinity DataIC50: 250nMAssay Description:Inhibition of USP2 (unknown origin) incubated for 20 mins followed by substrate addition by fluorescence based high-throughput assayMore data for this Ligand-Target Pair
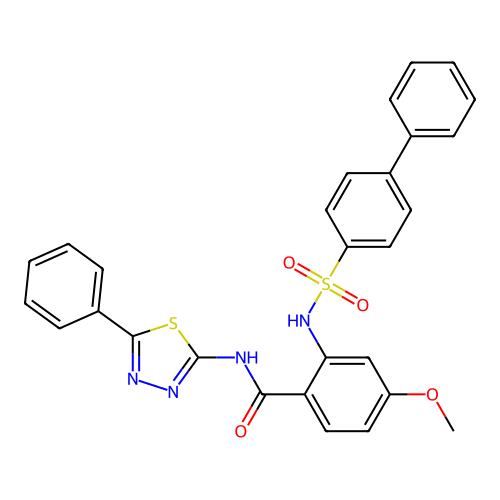
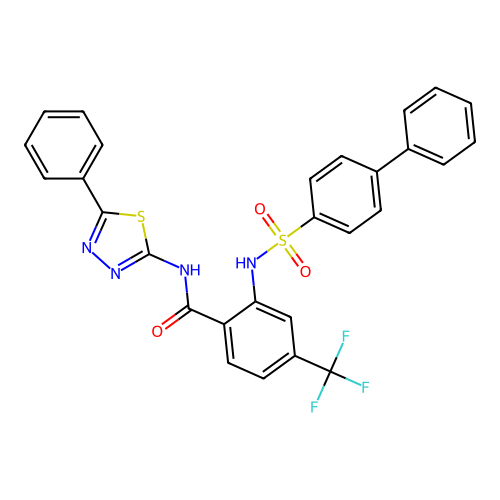
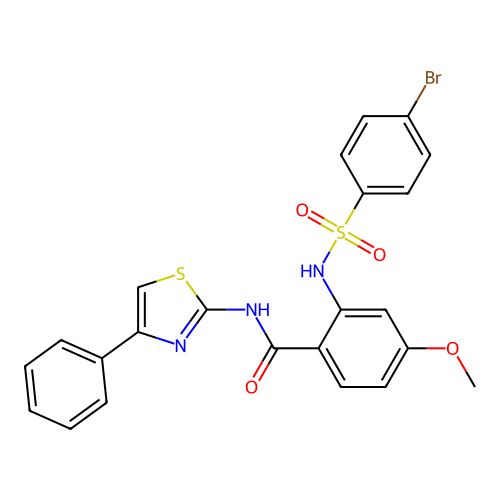
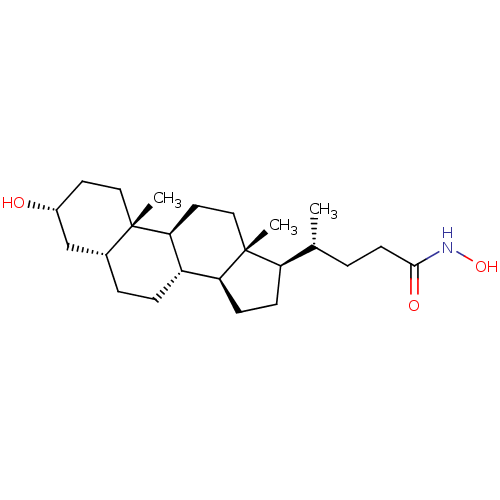
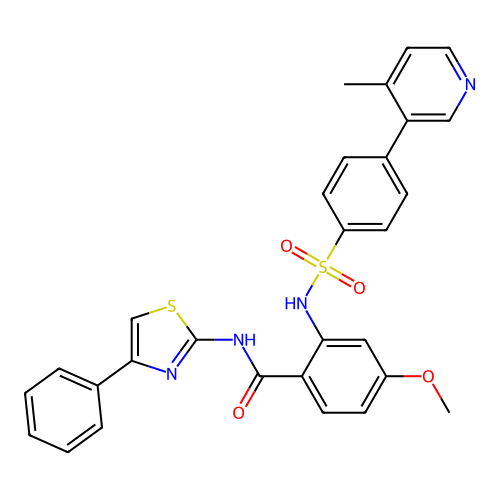
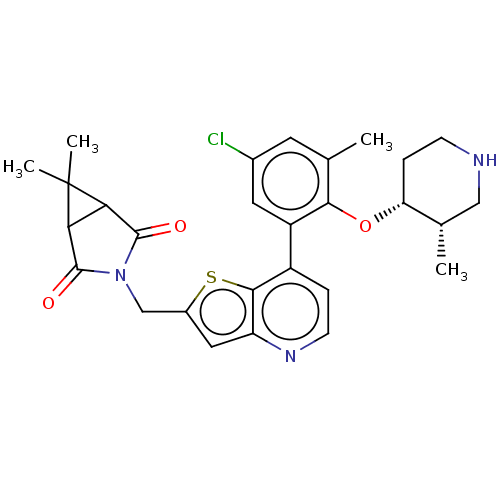
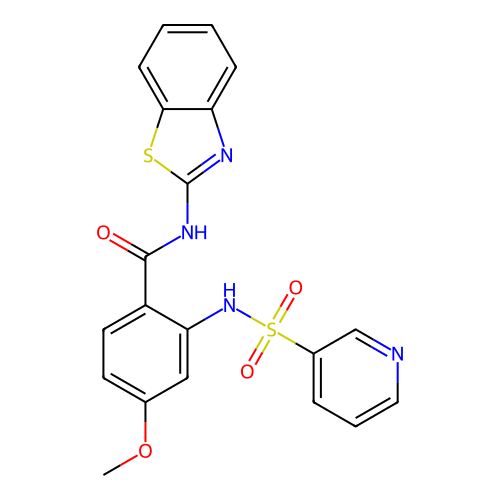
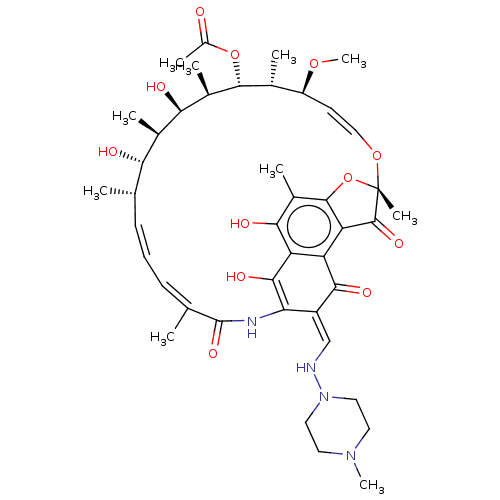
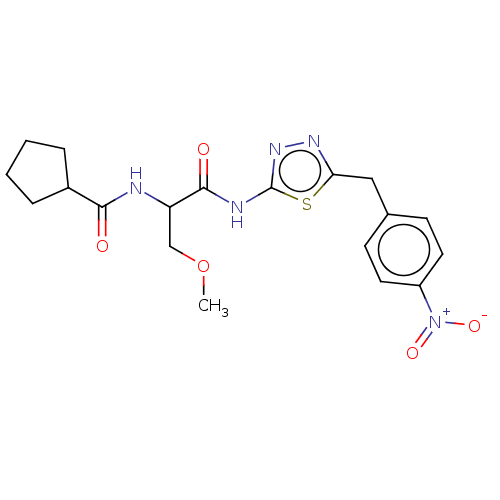
TargetUbiquitin carboxyl-terminal hydrolase 2(Human)
China Pharmaceutical University
Curated by ChEMBL
China Pharmaceutical University
Curated by ChEMBL
Affinity DataIC50: 800nMAssay Description:Inhibition of USP2 (267 to 599 residues) (unknown origin) expressed in Escherichia coli BL21 (DE3) cells using ubiquitin-rhodamine-110 as substrate p...More data for this Ligand-Target Pair
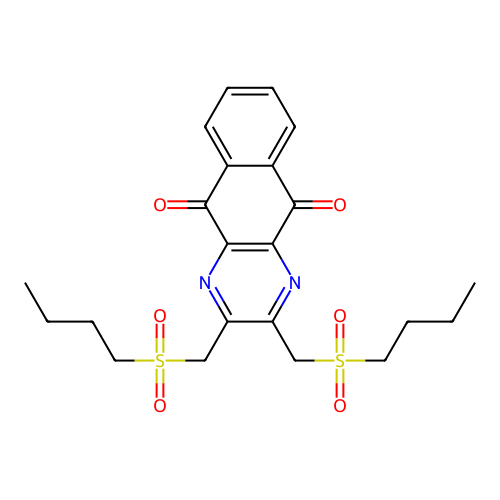
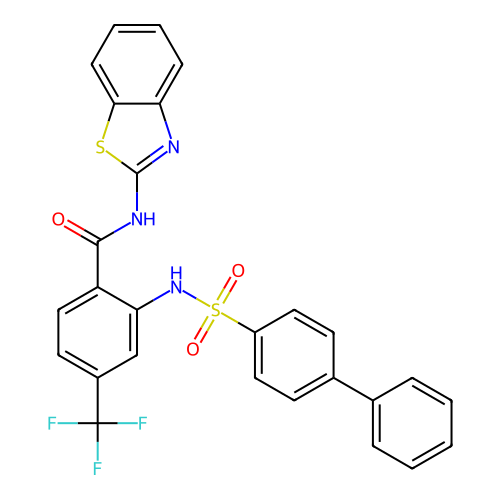
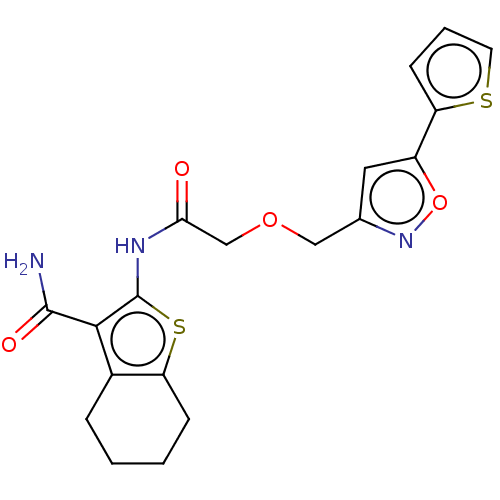
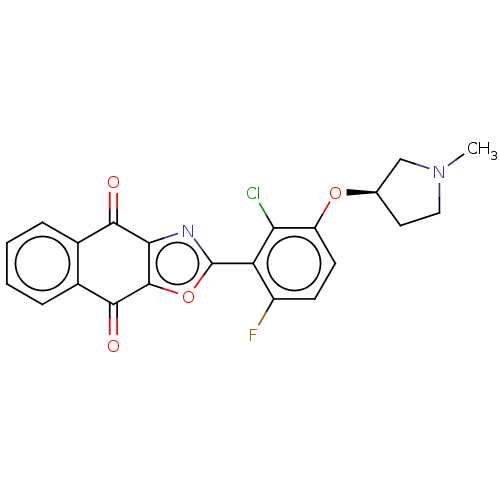
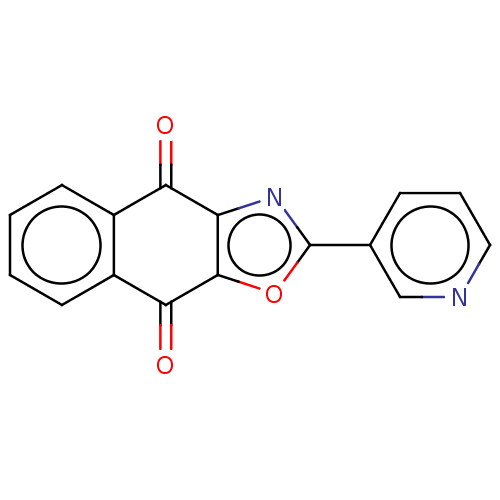
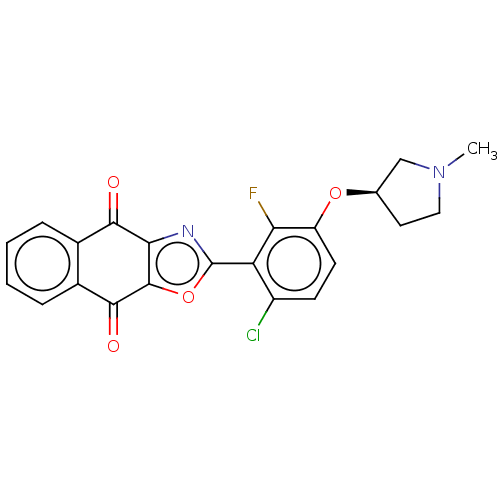
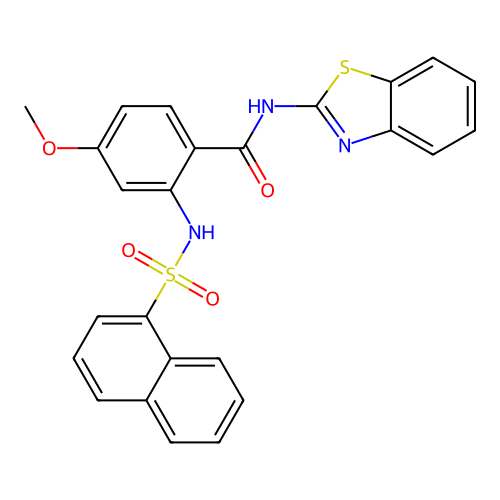
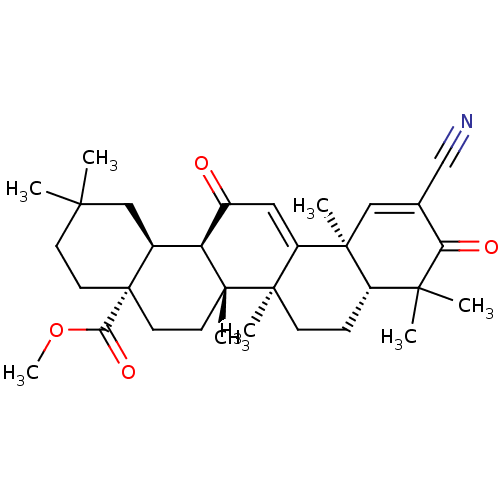
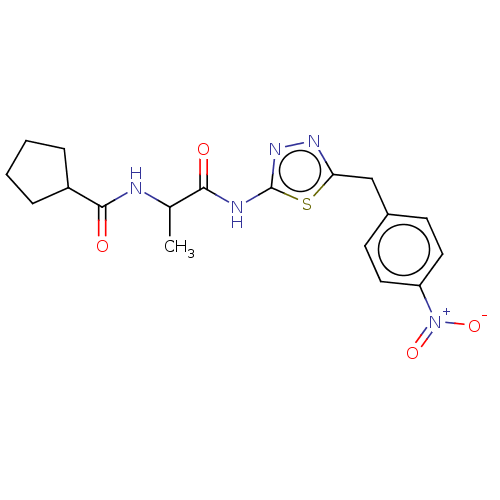
TargetUbiquitin carboxyl-terminal hydrolase 2(Human)
China Pharmaceutical University
Curated by ChEMBL
China Pharmaceutical University
Curated by ChEMBL
Affinity DataIC50: 800nMAssay Description:Inhibition of USP2 (unknown origin) using Di-Ub IQF K48-4 as substrate preincubated for 10 mins followed by substrate addition by high-throughput ass...More data for this Ligand-Target Pair
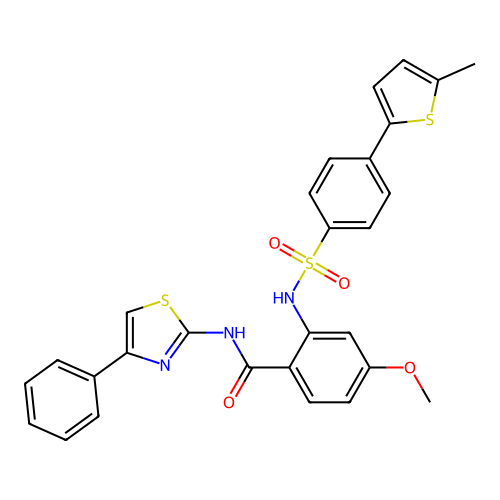
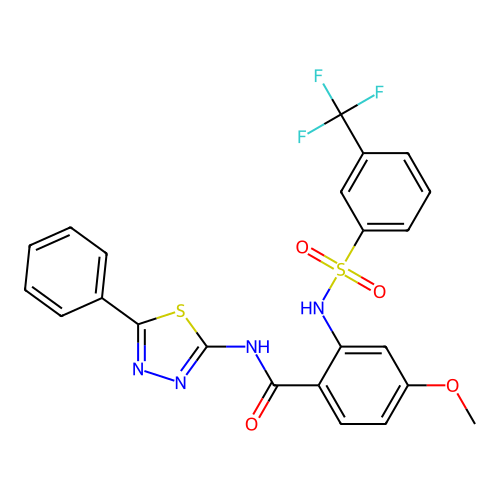
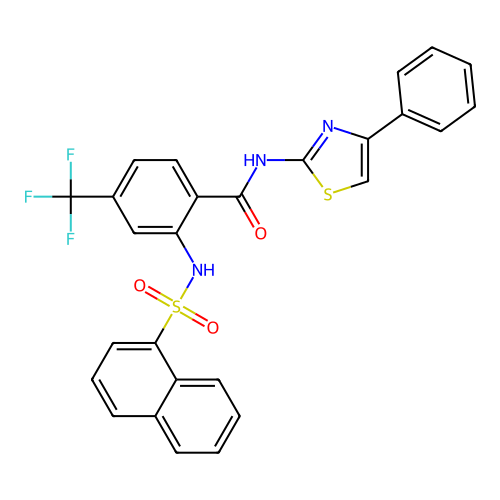
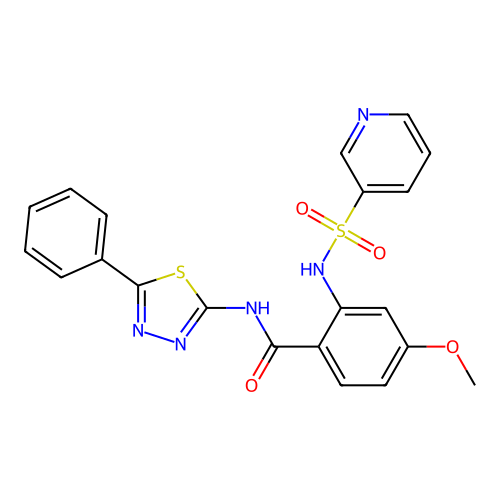
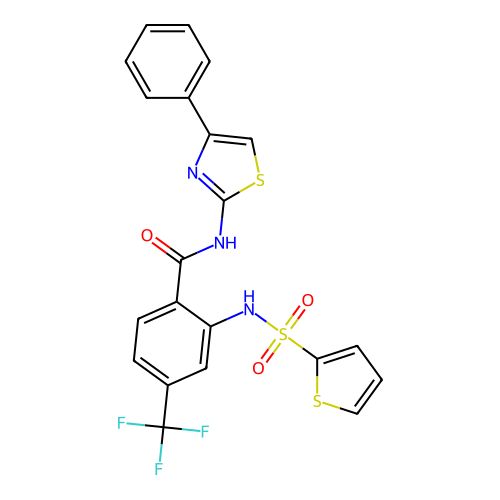
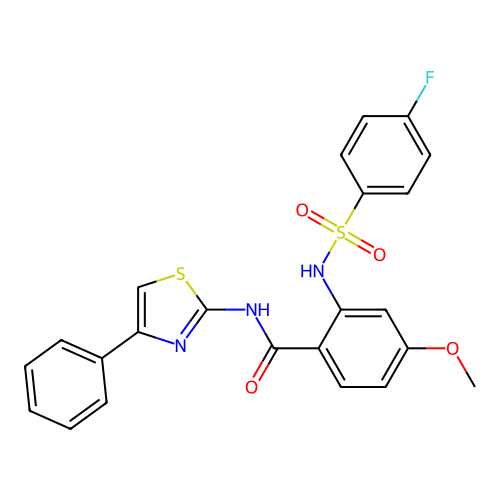
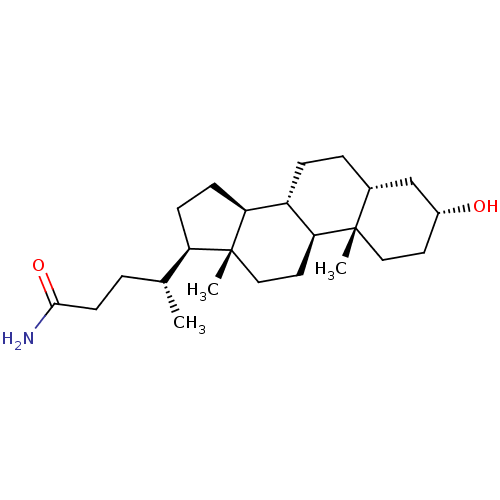
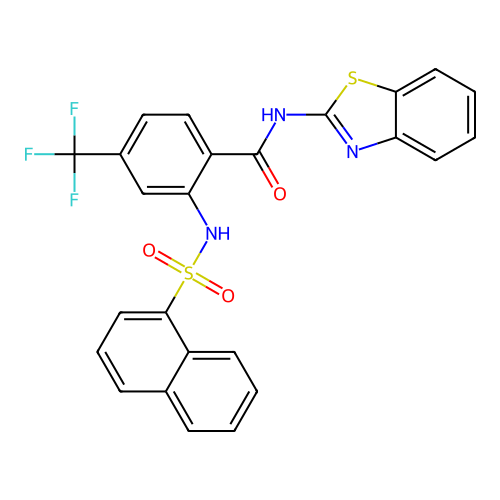
TargetUbiquitin carboxyl-terminal hydrolase 2(Human)
China Pharmaceutical University
Curated by ChEMBL
China Pharmaceutical University
Curated by ChEMBL
Affinity DataIC50: 890nMAssay Description:Inhibition of USP2 (267 to 599 residues) (unknown origin) expressed in Escherichia coli BL21 (DE3) cells using ubiquitin-rhodamine-110 as substrate p...More data for this Ligand-Target Pair
TargetUbiquitin carboxyl-terminal hydrolase 2(Human)
China Pharmaceutical University
Curated by ChEMBL
China Pharmaceutical University
Curated by ChEMBL
Affinity DataIC50: 1.10E+3nMAssay Description:Inhibition of USP2 (unknown origin) using Di-Ub IQF K48-4 as substrate preincubated for 10 mins followed by substrate addition by high-throughput ass...More data for this Ligand-Target Pair
TargetUbiquitin carboxyl-terminal hydrolase 2(Human)
China Pharmaceutical University
Curated by ChEMBL
China Pharmaceutical University
Curated by ChEMBL
Affinity DataIC50: 1.10E+3nMAssay Description:Inhibition of human USP2 using Di-Ub IQF as substrate incubated for 15 mins by K63-2 assayMore data for this Ligand-Target Pair
TargetUbiquitin carboxyl-terminal hydrolase 2(Human)
China Pharmaceutical University
Curated by ChEMBL
China Pharmaceutical University
Curated by ChEMBL
Affinity DataIC50: 1.30E+3nMAssay Description:Inhibition of USP2 (267 to 599 residues) (unknown origin) expressed in Escherichia coli BL21 (DE3) cells using ubiquitin-rhodamine-110 as substrate p...More data for this Ligand-Target Pair
TargetUbiquitin carboxyl-terminal hydrolase 2(Human)
China Pharmaceutical University
Curated by ChEMBL
China Pharmaceutical University
Curated by ChEMBL
Affinity DataIC50: 1.40E+3nMAssay Description:Inhibition of USP2 (267 to 599 residues) (unknown origin) expressed in Escherichia coli BL21 (DE3) cells using ubiquitin-rhodamine-110 as substrate p...More data for this Ligand-Target Pair
TargetUbiquitin carboxyl-terminal hydrolase 2(Human)
China Pharmaceutical University
Curated by ChEMBL
China Pharmaceutical University
Curated by ChEMBL
Affinity DataIC50: 1.50E+3nMAssay Description:Inhibition of USP2 (267 to 599 residues) (unknown origin) expressed in Escherichia coli BL21 (DE3) cells using ubiquitin-rhodamine-110 as substrate p...More data for this Ligand-Target Pair
TargetUbiquitin carboxyl-terminal hydrolase 2(Human)
China Pharmaceutical University
Curated by ChEMBL
China Pharmaceutical University
Curated by ChEMBL
Affinity DataIC50: 1.60E+3nMAssay Description:Inhibition of USP2 (267 to 599 residues) (unknown origin) expressed in Escherichia coli BL21 (DE3) cells using ubiquitin-rhodamine-110 as substrate p...More data for this Ligand-Target Pair
TargetUbiquitin carboxyl-terminal hydrolase 2(Human)
China Pharmaceutical University
Curated by ChEMBL
China Pharmaceutical University
Curated by ChEMBL
Affinity DataIC50: 3.20E+3nMAssay Description:Inhibition of USP2 (267 to 599 residues) (unknown origin) expressed in Escherichia coli BL21 (DE3) cells using ubiquitin-rhodamine-110 as substrate p...More data for this Ligand-Target Pair
TargetUbiquitin carboxyl-terminal hydrolase 2(Human)
China Pharmaceutical University
Curated by ChEMBL
China Pharmaceutical University
Curated by ChEMBL
Affinity DataIC50: 3.30E+3nMAssay Description:Inhibition of human USP2a (258 to 605 residues) expressed in Escherichia coli BL21(DE3) using Ub-AMC as substrate incubated for 30 mins measured at 3...More data for this Ligand-Target Pair
Affinity DataIC50: 3.30E+3nMAssay Description:The assays were performed using Infinite 200 PRO - Tecan plate reader and 96-well, black Greiner microplates in a 100 ml reaction volume. The assays ...More data for this Ligand-Target Pair
TargetUbiquitin carboxyl-terminal hydrolase 2(Human)
China Pharmaceutical University
Curated by ChEMBL
China Pharmaceutical University
Curated by ChEMBL
Affinity DataIC50: 3.30E+3nMAssay Description:Inhibition of USP2 (267 to 599 residues) (unknown origin) expressed in Escherichia coli BL21 (DE3) cells using ubiquitin-rhodamine-110 as substrate p...More data for this Ligand-Target Pair
TargetUbiquitin carboxyl-terminal hydrolase 2(Human)
China Pharmaceutical University
Curated by ChEMBL
China Pharmaceutical University
Curated by ChEMBL
Affinity DataIC50: 3.40E+3nMAssay Description:Inhibition of USP2 (267 to 599 residues) (unknown origin) expressed in Escherichia coli BL21 (DE3) cells using ubiquitin-rhodamine-110 as substrate p...More data for this Ligand-Target Pair
TargetUbiquitin carboxyl-terminal hydrolase 2(Human)
China Pharmaceutical University
Curated by ChEMBL
China Pharmaceutical University
Curated by ChEMBL
Affinity DataIC50: 3.60E+3nMAssay Description:Inhibition of USP2 (267 to 599 residues) (unknown origin) expressed in Escherichia coli BL21 (DE3) cells using ubiquitin-rhodamine-110 as substrate p...More data for this Ligand-Target Pair
Affinity DataIC50: 3.70E+3nMAssay Description:The assays were performed using Infinite 200 PRO - Tecan plate reader and 96-well, black Greiner microplates in a 100 ml reaction volume. The assays ...More data for this Ligand-Target Pair
TargetUbiquitin carboxyl-terminal hydrolase 2(Human)
China Pharmaceutical University
Curated by ChEMBL
China Pharmaceutical University
Curated by ChEMBL
Affinity DataIC50: 3.80E+3nMAssay Description:Inhibition of USP2 (unknown origin) using Ub-Rho measured after 0.5 to 3 hrsMore data for this Ligand-Target Pair
TargetUbiquitin carboxyl-terminal hydrolase 2(Human)
China Pharmaceutical University
Curated by ChEMBL
China Pharmaceutical University
Curated by ChEMBL
Affinity DataIC50: 3.80E+3nMAssay Description:Inhibition of USP2 (267 to 599 residues) (unknown origin) expressed in Escherichia coli BL21 (DE3) cells using ubiquitin-rhodamine-110 as substrate p...More data for this Ligand-Target Pair
TargetUbiquitin carboxyl-terminal hydrolase 2(Human)
China Pharmaceutical University
Curated by ChEMBL
China Pharmaceutical University
Curated by ChEMBL
Affinity DataIC50: 4.40E+3nMAssay Description:Inhibition of USP2 (267 to 599 residues) (unknown origin) expressed in Escherichia coli BL21 (DE3) cells using ubiquitin-rhodamine-110 as substrate p...More data for this Ligand-Target Pair
TargetUbiquitin carboxyl-terminal hydrolase 2(Human)
China Pharmaceutical University
Curated by ChEMBL
China Pharmaceutical University
Curated by ChEMBL
Affinity DataIC50: 4.60E+3nMAssay Description:Inhibition of USP2 (267 to 599 residues) (unknown origin) expressed in Escherichia coli BL21 (DE3) cells using ubiquitin-rhodamine-110 as substrate p...More data for this Ligand-Target Pair
TargetUbiquitin carboxyl-terminal hydrolase 2(Human)
China Pharmaceutical University
Curated by ChEMBL
China Pharmaceutical University
Curated by ChEMBL
Affinity DataIC50: 5.10E+3nMAssay Description:Inhibition of USP2 (unknown origin) using Ub-Rho measured after 0.5 to 3 hrsMore data for this Ligand-Target Pair
TargetUbiquitin carboxyl-terminal hydrolase 2(Human)
China Pharmaceutical University
Curated by ChEMBL
China Pharmaceutical University
Curated by ChEMBL
Affinity DataIC50: 5.50E+3nMAssay Description:Inhibition of USP2 (267 to 599 residues) (unknown origin) expressed in Escherichia coli BL21 (DE3) cells using ubiquitin-rhodamine-110 as substrate p...More data for this Ligand-Target Pair
TargetUbiquitin carboxyl-terminal hydrolase 2(Human)
China Pharmaceutical University
Curated by ChEMBL
China Pharmaceutical University
Curated by ChEMBL
Affinity DataIC50: 5.50E+3nMAssay Description:Inhibition of USP2 (267 to 599 residues) (unknown origin) expressed in Escherichia coli BL21 (DE3) cells using ubiquitin-rhodamine-110 as substrate p...More data for this Ligand-Target Pair
TargetUbiquitin carboxyl-terminal hydrolase 2(Human)
China Pharmaceutical University
Curated by ChEMBL
China Pharmaceutical University
Curated by ChEMBL
Affinity DataIC50: 5.80E+3nMAssay Description:Inhibition of human USP2A (258 to 605 residues) expressed in Escherichia coli BL21 (DE3) using Ub-AMC as substrate preincubated for 30 mins followed ...More data for this Ligand-Target Pair
Affinity DataIC50: 5.80E+3nMAssay Description:The assays were performed using Infinite 200 PRO - Tecan plate reader and 96-well, black Greiner microplates in a 100 ml reaction volume. The assays ...More data for this Ligand-Target Pair
TargetUbiquitin carboxyl-terminal hydrolase 2(Human)
China Pharmaceutical University
Curated by ChEMBL
China Pharmaceutical University
Curated by ChEMBL
Affinity DataIC50: 5.90E+3nMAssay Description:Inhibition of USP2 (267 to 599 residues) (unknown origin) expressed in Escherichia coli BL21 (DE3) cells using ubiquitin-rhodamine-110 as substrate p...More data for this Ligand-Target Pair
TargetUbiquitin carboxyl-terminal hydrolase 2(Human)
China Pharmaceutical University
Curated by ChEMBL
China Pharmaceutical University
Curated by ChEMBL
Affinity DataIC50: 6.20E+3nMAssay Description:Inhibition of USP2 (unknown origin) using Ub-Rho measured after 0.5 to 3 hrsMore data for this Ligand-Target Pair
TargetUbiquitin carboxyl-terminal hydrolase 2(Human)
China Pharmaceutical University
Curated by ChEMBL
China Pharmaceutical University
Curated by ChEMBL
Affinity DataIC50: 6.40E+3nMAssay Description:Inhibition of USP2 (267 to 599 residues) (unknown origin) expressed in Escherichia coli BL21 (DE3) cells using ubiquitin-rhodamine-110 as substrate p...More data for this Ligand-Target Pair
TargetUbiquitin carboxyl-terminal hydrolase 2(Human)
China Pharmaceutical University
Curated by ChEMBL
China Pharmaceutical University
Curated by ChEMBL
Affinity DataEC50: 7.20E+3nMAssay Description:Inhibition of USP2 (unknown origin)More data for this Ligand-Target Pair
Affinity DataIC50: 7.40E+3nMAssay Description:The assays were performed using Infinite 200 PRO - Tecan plate reader and 96-well, black Greiner microplates in a 100 ml reaction volume. The assays ...More data for this Ligand-Target Pair
TargetUbiquitin carboxyl-terminal hydrolase 2(Human)
China Pharmaceutical University
Curated by ChEMBL
China Pharmaceutical University
Curated by ChEMBL
Affinity DataIC50: 8.70E+3nMAssay Description:Inhibition of human USP2a (258 to 605 residues) expressed in Escherichia coli BL21(DE3) using Ub-AMC as substrate incubated for 30 mins measured at 3...More data for this Ligand-Target Pair
TargetUbiquitin carboxyl-terminal hydrolase 2(Human)
China Pharmaceutical University
Curated by ChEMBL
China Pharmaceutical University
Curated by ChEMBL
Affinity DataIC50: 9.70E+3nMAssay Description:Inhibition of human USP2A (258 to 605 residues) expressed in Escherichia coli BL21 (DE3) using Ub-AMC as substrate pre-incubated for 30 mins followed...More data for this Ligand-Target Pair
TargetUbiquitin carboxyl-terminal hydrolase 2(Human)
China Pharmaceutical University
Curated by ChEMBL
China Pharmaceutical University
Curated by ChEMBL
Affinity DataIC50: 9.70E+3nMAssay Description:Inhibition of USP2 (unknown origin) using Ub-AMC substrateMore data for this Ligand-Target Pair
Affinity DataIC50: 9.70E+3nMAssay Description:The assays were performed using Infinite 200 PRO - Tecan plate reader and 96-well, black Greiner microplates in a 100 ml reaction volume. The assays ...More data for this Ligand-Target Pair
TargetUbiquitin carboxyl-terminal hydrolase 2(Human)
China Pharmaceutical University
Curated by ChEMBL
China Pharmaceutical University
Curated by ChEMBL
Affinity DataIC50: 1.00E+4nMAssay Description:Inhibition of human GST-tagged USP2 isoform 4 expressed in Escherichia coli assessed as cleavage of Ubiquitin-Rhodamine110-glycine to Ubiquitin and R...More data for this Ligand-Target Pair
TargetUbiquitin carboxyl-terminal hydrolase 2(Human)
China Pharmaceutical University
Curated by ChEMBL
China Pharmaceutical University
Curated by ChEMBL
Affinity DataIC50: 1.00E+4nMAssay Description:Inhibition of USP2 (unknown origin) using Ub-Rho measured after 0.5 to 3 hrsMore data for this Ligand-Target Pair
TargetUbiquitin carboxyl-terminal hydrolase 2(Human)
China Pharmaceutical University
Curated by ChEMBL
China Pharmaceutical University
Curated by ChEMBL
Affinity DataIC50: 1.05E+4nMAssay Description:Inhibition of USP2 (267 to 599 residues) (unknown origin) expressed in Escherichia coli BL21 (DE3) cells using ubiquitin-rhodamine-110 as substrate p...More data for this Ligand-Target Pair
TargetUbiquitin carboxyl-terminal hydrolase 2(Human)
China Pharmaceutical University
Curated by ChEMBL
China Pharmaceutical University
Curated by ChEMBL
Affinity DataIC50: 1.33E+4nMAssay Description:Inhibition of USP2 (267 to 599 residues) (unknown origin) expressed in Escherichia coli BL21 (DE3) cells using ubiquitin-rhodamine-110 as substrate p...More data for this Ligand-Target Pair
Affinity DataIC50: 1.39E+4nMAssay Description:The assays were performed using Infinite 200 PRO - Tecan plate reader and 96-well, black Greiner microplates in a 100 ml reaction volume. The assays ...More data for this Ligand-Target Pair
TargetUbiquitin carboxyl-terminal hydrolase 2(Human)
China Pharmaceutical University
Curated by ChEMBL
China Pharmaceutical University
Curated by ChEMBL
Affinity DataIC50: 1.45E+4nMAssay Description:Inhibition of USP2 (267 to 599 residues) (unknown origin) expressed in Escherichia coli BL21 (DE3) cells using ubiquitin-rhodamine-110 as substrate p...More data for this Ligand-Target Pair
TargetUbiquitin carboxyl-terminal hydrolase 2(Human)
China Pharmaceutical University
Curated by ChEMBL
China Pharmaceutical University
Curated by ChEMBL
Affinity DataIC50: 1.54E+4nMAssay Description:Inhibition of USP2 (267 to 599 residues) (unknown origin) expressed in Escherichia coli BL21 (DE3) cells using ubiquitin-rhodamine-110 as substrate p...More data for this Ligand-Target Pair
Affinity DataIC50: 1.67E+4nMAssay Description:The assays were performed using Infinite 200 PRO - Tecan plate reader and 96-well, black Greiner microplates in a 100 ml reaction volume. The assays ...More data for this Ligand-Target Pair
TargetUbiquitin carboxyl-terminal hydrolase 2(Human)
China Pharmaceutical University
Curated by ChEMBL
China Pharmaceutical University
Curated by ChEMBL
Affinity DataIC50: 1.82E+4nMpH: 7.5 T: 2°CAssay Description:Ubiquitin-7-amino-4-methylcoumarin (Ub-AMC) was generated as previously described.(7) The enzymatic reaction was conducted in fluorometric assay buff...More data for this Ligand-Target Pair
TargetUbiquitin carboxyl-terminal hydrolase 2(Human)
China Pharmaceutical University
Curated by ChEMBL
China Pharmaceutical University
Curated by ChEMBL
Affinity DataIC50: 2.12E+4nMpH: 7.5 T: 2°CAssay Description:Ubiquitin-7-amino-4-methylcoumarin (Ub-AMC) was generated as previously described.(7) The enzymatic reaction was conducted in fluorometric assay buff...More data for this Ligand-Target Pair
TargetUbiquitin carboxyl-terminal hydrolase 2(Human)
China Pharmaceutical University
Curated by ChEMBL
China Pharmaceutical University
Curated by ChEMBL
Affinity DataIC50: 2.20E+4nMAssay Description:Inhibition of USP2 (unknown origin)More data for this Ligand-Target Pair
TargetUbiquitin carboxyl-terminal hydrolase 2(Human)
China Pharmaceutical University
Curated by ChEMBL
China Pharmaceutical University
Curated by ChEMBL
Affinity DataIC50: 2.23E+4nMAssay Description:Inhibition of USP2 (267 to 599 residues) (unknown origin) expressed in Escherichia coli BL21 (DE3) cells using ubiquitin-rhodamine-110 as substrate p...More data for this Ligand-Target Pair
TargetUbiquitin carboxyl-terminal hydrolase 2(Human)
China Pharmaceutical University
Curated by ChEMBL
China Pharmaceutical University
Curated by ChEMBL
Affinity DataIC50: 2.50E+4nMAssay Description:Inhibition of human His-tagged USP2 (259 to 605 residues) using ubiquitin rhodamine 110 110 as substrate preincubated for 15 to 20 mins followed by e...More data for this Ligand-Target Pair
TargetUbiquitin carboxyl-terminal hydrolase 2(Human)
China Pharmaceutical University
Curated by ChEMBL
China Pharmaceutical University
Curated by ChEMBL
Affinity DataIC50: 2.50E+4nMAssay Description:Inhibition of human His-tagged USP2 (259 to 605 residues) using ubiquitin rhodamine 110 110 as substrate preincubated for 15 to 20 mins followed by e...More data for this Ligand-Target Pair
TargetUbiquitin carboxyl-terminal hydrolase 2(Human)
China Pharmaceutical University
Curated by ChEMBL
China Pharmaceutical University
Curated by ChEMBL
Affinity DataIC50: 2.50E+4nMAssay Description:Inhibition of human His-tagged USP2 (259 to 605 residues) using ubiquitin rhodamine 110 110 as substrate preincubated for 15 to 20 mins followed by e...More data for this Ligand-Target Pair