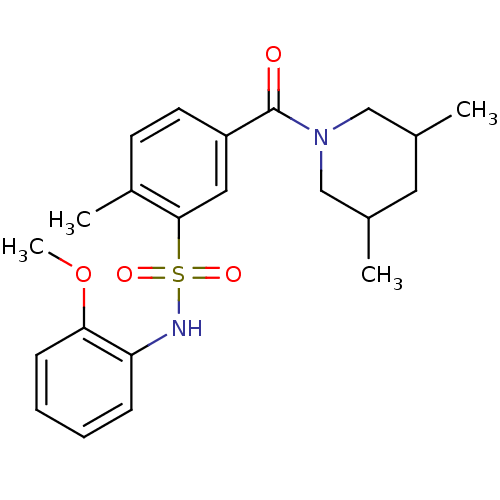
BDBM72021 5-(3,5-dimethylpiperidin-1-yl)carbonyl-N-(2-methoxyphenyl)-2-methyl-benzenesulfonamide::5-(3,5-dimethylpiperidine-1-carbonyl)-N-(2-methoxyphenyl)-2-methyl-benzenesulfonamide::5-(3,5-dimethylpiperidine-1-carbonyl)-N-(2-methoxyphenyl)-2-methylbenzenesulfonamide::5-[(3,5-dimethyl-1-piperidinyl)-oxomethyl]-N-(2-methoxyphenyl)-2-methylbenzenesulfonamide::MLS000536447::SMR000151379::cid_5069540
SMILES COc1ccccc1NS(=O)(=O)c1cc(ccc1C)C(=O)N1CC(C)CC(C)C1
InChI Key InChIKey=VFUQTHVFXSUIAI-UHFFFAOYSA-N
Data 1 EC50
Activity Spreadsheet -- Enzyme Inhibition Constant Data from BindingDB
 Found 2 hits for monomerid = 72021
Found 2 hits for monomerid = 72021
Affinity DataKi: 1.57E+3nMAssay Description:Keywords: Heat Shock Factor-1 (HSF-1), Stress Response, MG132, NIH3T3, Luminescence Assay Overview: Modified NIH3T3, transformed to express firefly...More data for this Ligand-Target Pair
TargetPotassium voltage-gated channel subfamily H member 2(Human)
California Institute Of Technology
US Patent
California Institute Of Technology
US Patent
Affinity DataKi: 7.21E+3nMAssay Description:Keywords: Heat Shock Factor-1 (HSF-1), Stress Response, MG132, NIH3T3, Luminescence Assay Overview: Modified NIH3T3, transformed to express firefly...More data for this Ligand-Target Pair