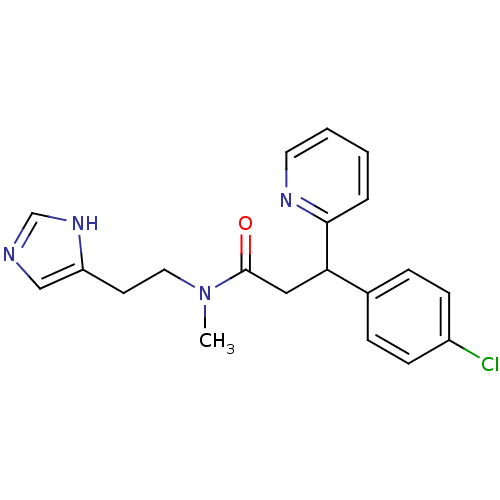
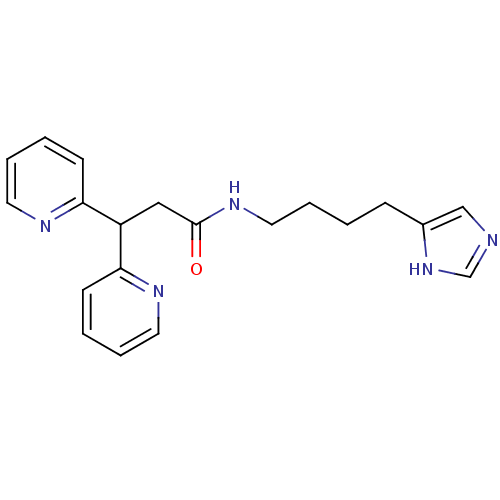
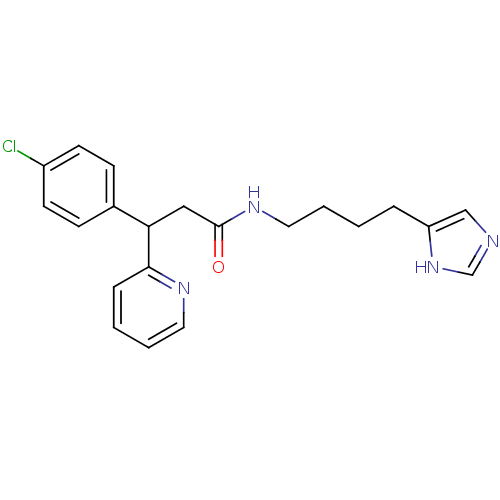
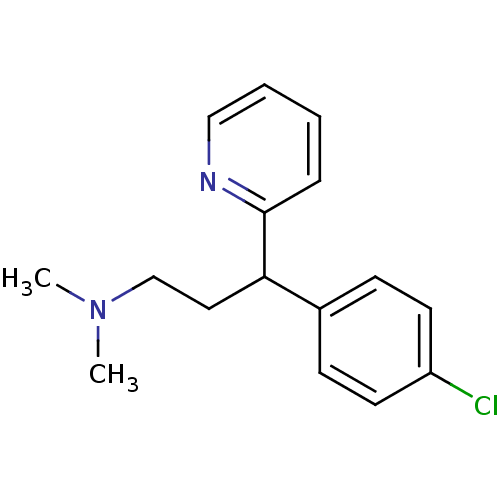
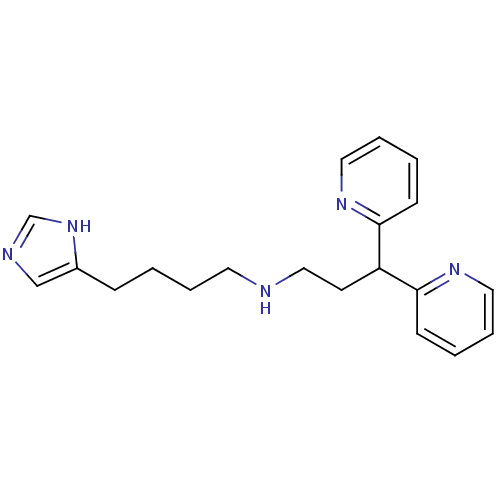
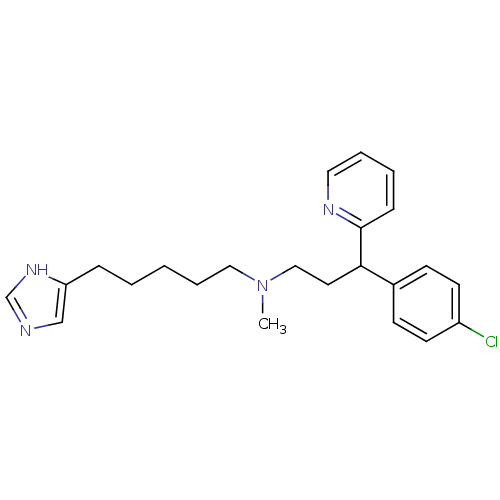
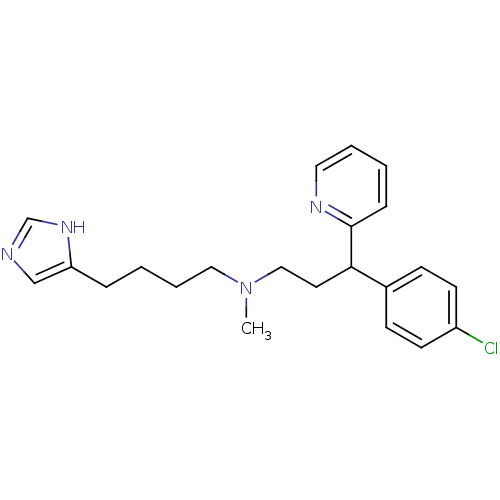
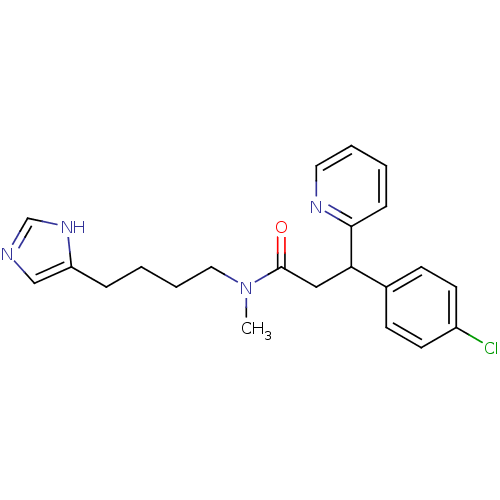
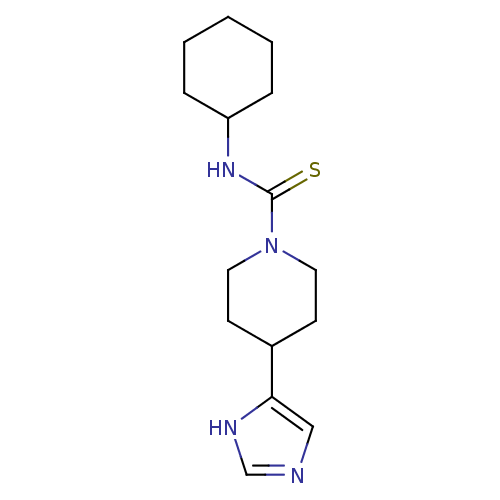
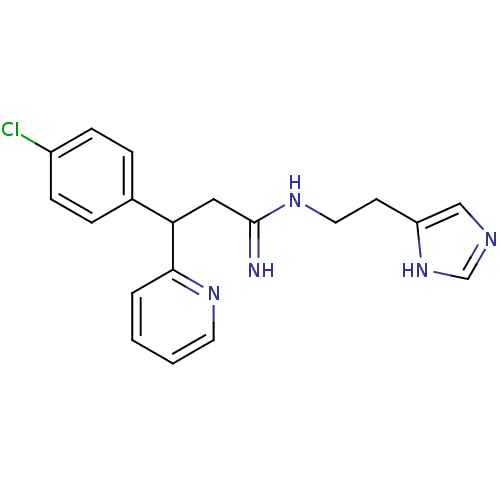
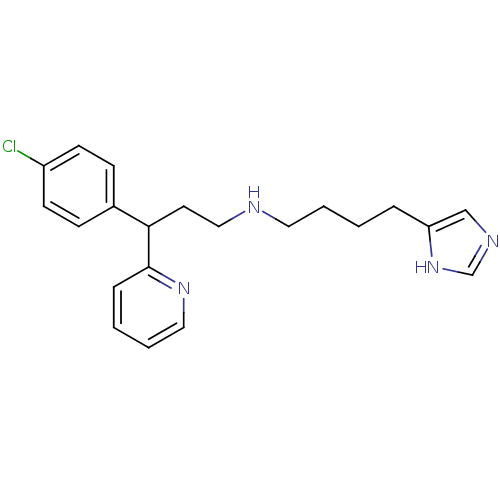
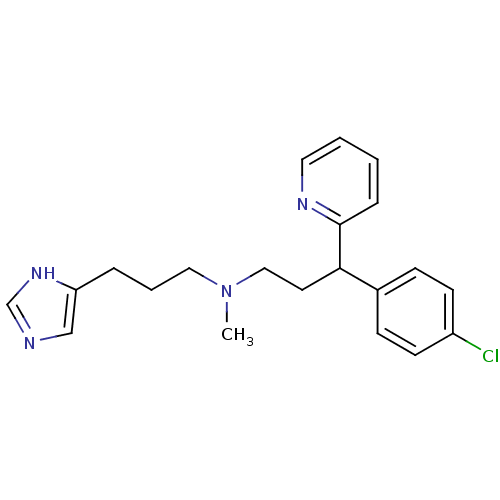
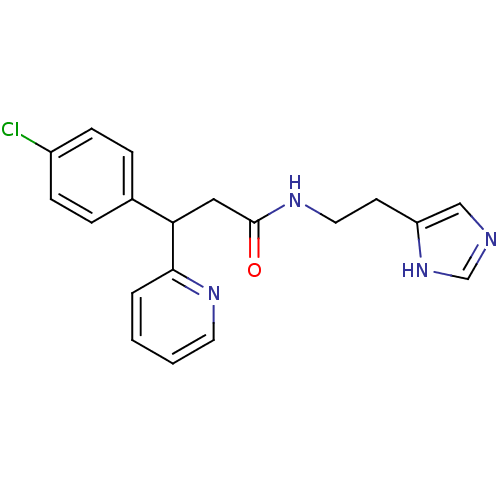
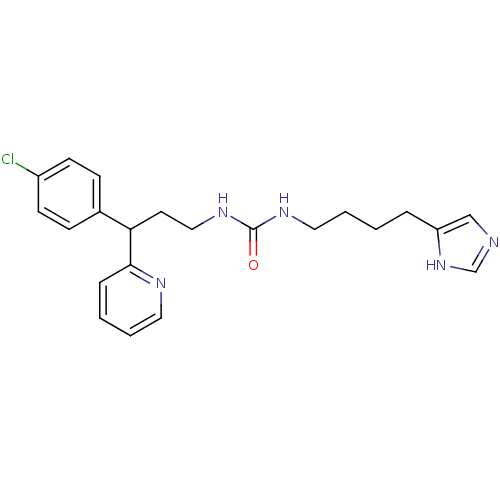
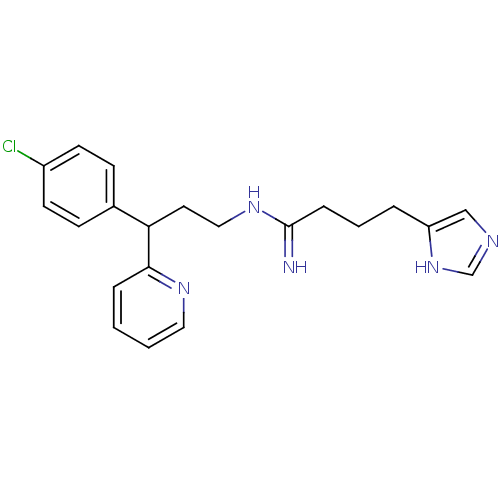
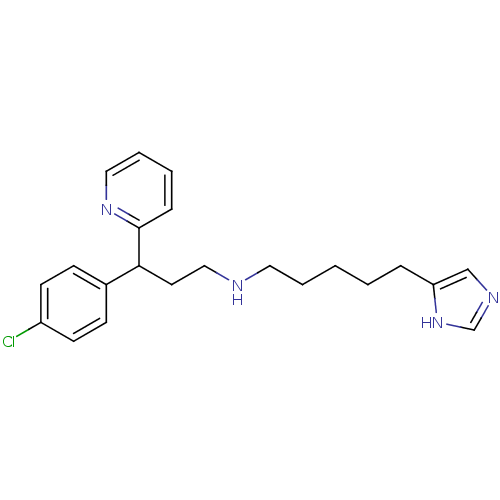
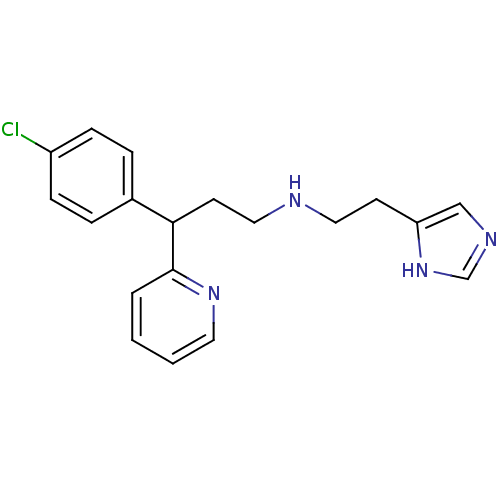
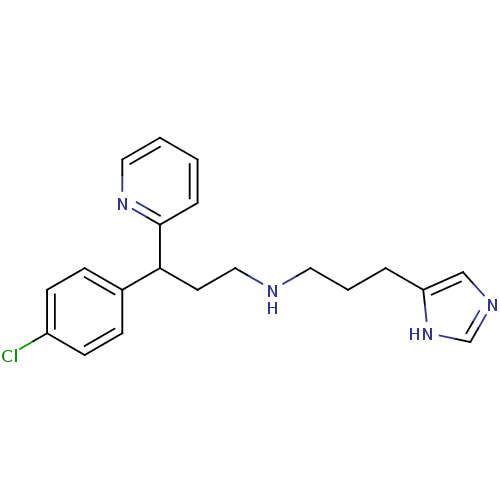
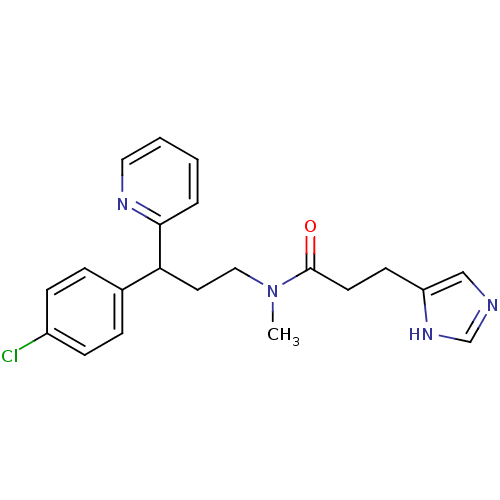
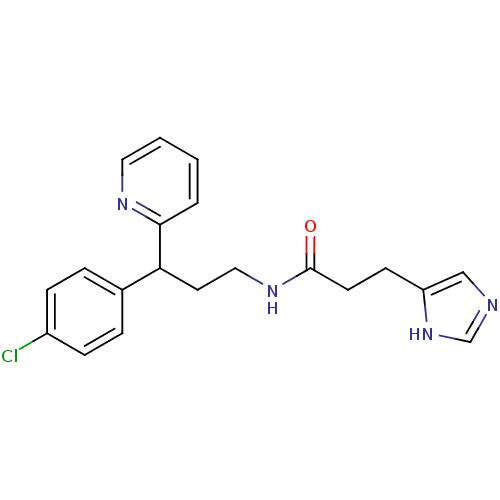
Report error Found 28 Enz. Inhib. hit(s) with all data for entry = 50013301
Affinity DataKi: 1nMAssay Description:Binding affinity against H3 receptor in guinea pig brain using [3H]N-alpha-methyl histamine as radioligandMore data for this Ligand-Target Pair
Affinity DataKi: 1nMAssay Description:Binding affinity against H3 receptor in guinea pig brain using [3H]N-alpha-methyl histamine as radioligandMore data for this Ligand-Target Pair
Affinity DataKi: 1nMAssay Description:Binding affinity against H3 receptor in guinea pig brain using [3H]N-alpha-methyl histamine as radioligandMore data for this Ligand-Target Pair
Affinity DataKi: 2nMAssay Description:Ability to displace [3H]pyrilamine from histamine H1 receptor in male Sprague-Dawley rat brain membranesMore data for this Ligand-Target Pair
Affinity DataKi: 4nMAssay Description:Binding affinity against H3 receptor in guinea pig brain using [3H]N-alpha-methyl histamine as radioligandMore data for this Ligand-Target Pair
Affinity DataKi: 6nMAssay Description:Ability to displace [3H]pyrilamine from histamine H1 receptor in male Sprague-Dawley rat brain membranesMore data for this Ligand-Target Pair
Affinity DataKi: 7nMAssay Description:Ability to displace [3H]pyrilamine from histamine H1 receptor in male Sprague-Dawley rat brain membranesMore data for this Ligand-Target Pair
Affinity DataKi: 7nMAssay Description:Binding affinity against H3 receptor in guinea pig brain using [3H]N-alpha-methyl histamine as radioligandMore data for this Ligand-Target Pair
Affinity DataKi: 7.30nMAssay Description:Binding affinity against H3 receptor in guinea pig brain using [3H]N-alpha-methyl histamine as radioligandMore data for this Ligand-Target Pair
Affinity DataKi: 8nMAssay Description:Binding affinity against H3 receptor in guinea pig brain using [3H]N-alpha-methyl histamine as radioligandMore data for this Ligand-Target Pair
Affinity DataKi: 10nMAssay Description:Binding affinity against H3 receptor in guinea pig brain using [3H]N-alpha-methyl histamine as radioligandMore data for this Ligand-Target Pair
Affinity DataKi: 10nMAssay Description:Ability to displace [3H]pyrilamine from histamine H1 receptor in male Sprague-Dawley rat brain membranesMore data for this Ligand-Target Pair
Affinity DataKi: 15nMAssay Description:Binding affinity against H3 receptor in guinea pig brain using [3H]N-alpha-methyl histamine as radioligandMore data for this Ligand-Target Pair
Affinity DataKi: 16nMAssay Description:Binding affinity against H3 receptor in guinea pig brain using [3H]N-alpha-methyl histamine as radioligandMore data for this Ligand-Target Pair
Affinity DataKi: 23nMAssay Description:Binding affinity against H3 receptor in guinea pig brain using [3H]N-alpha-methyl histamine as radioligandMore data for this Ligand-Target Pair
Affinity DataKi: 29nMAssay Description:Binding affinity against H3 receptor in guinea pig brain using [3H]N-alpha-methyl histamine as radioligandMore data for this Ligand-Target Pair
Affinity DataKi: 33nMAssay Description:Binding affinity against H3 receptor in guinea pig brain using [3H]N-alpha-methyl histamine as radioligandMore data for this Ligand-Target Pair
Affinity DataKi: 39nMAssay Description:Ability to displace [3H]pyrilamine from histamine H1 receptor in male Sprague-Dawley rat brain membranesMore data for this Ligand-Target Pair
Affinity DataKi: 50nMAssay Description:Binding affinity against H3 receptor in guinea pig brain using [3H]N-alpha-methyl histamine as radioligandMore data for this Ligand-Target Pair
Affinity DataKi: 56nMAssay Description:Binding affinity against H3 receptor in guinea pig brain using [3H]N-alpha-methyl histamine as radioligandMore data for this Ligand-Target Pair
Affinity DataKi: 120nMAssay Description:Binding affinity against H3 receptor in guinea pig brain using [3H]N-alpha-methyl histamine as radioligandMore data for this Ligand-Target Pair
Affinity DataKi: 201nMAssay Description:Ability to displace [3H]pyrilamine from histamine H1 receptor in male Sprague-Dawley rat brain membranesMore data for this Ligand-Target Pair
Affinity DataKi: 240nMAssay Description:Binding affinity against H3 receptor in guinea pig brain using [3H]N-alpha-methyl histamine as radioligandMore data for this Ligand-Target Pair
Affinity DataKi: 254nMAssay Description:Ability to displace [3H]pyrilamine from histamine H1 receptor in male Sprague-Dawley rat brain membranesMore data for this Ligand-Target Pair
Affinity DataKi: 260nMAssay Description:Binding affinity against H3 receptor in guinea pig brain using [3H]N-alpha-methyl histamine as radioligandMore data for this Ligand-Target Pair
Affinity DataKi: 510nMAssay Description:Binding affinity against H3 receptor in guinea pig brain using [3H]N-alpha-methyl histamine as radioligandMore data for this Ligand-Target Pair
Affinity DataKi: 600nMAssay Description:Ability to displace [3H]pyrilamine from histamine H1 receptor in male Sprague-Dawley rat brain membranesMore data for this Ligand-Target Pair
Affinity DataKi: 920nMAssay Description:Ability to displace [3H]pyrilamine from histamine H1 receptor in male Sprague-Dawley rat brain membranesMore data for this Ligand-Target Pair