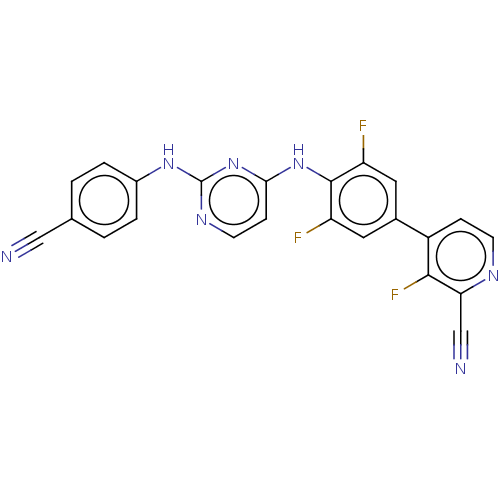
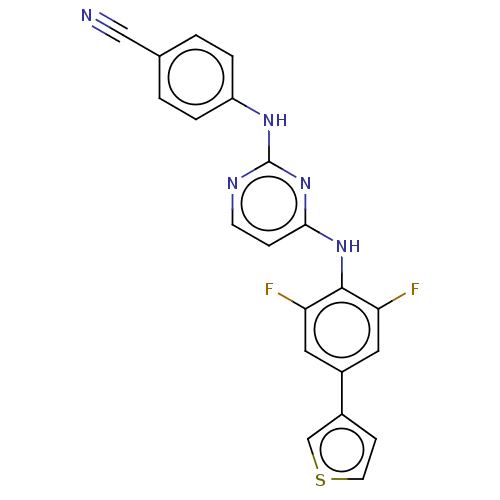
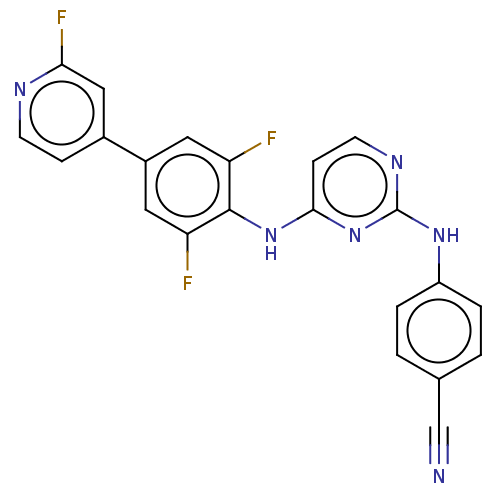
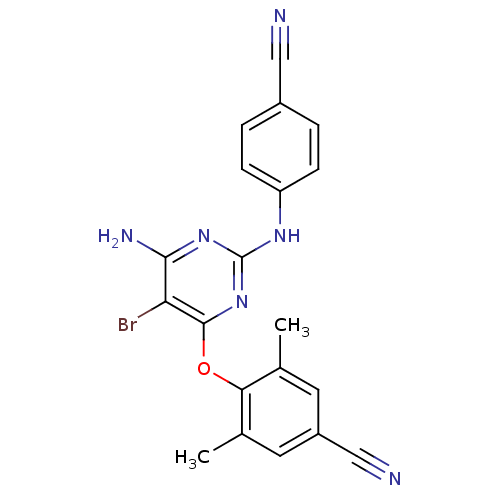
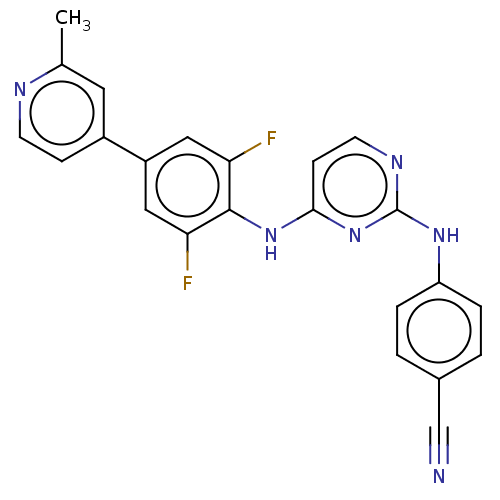
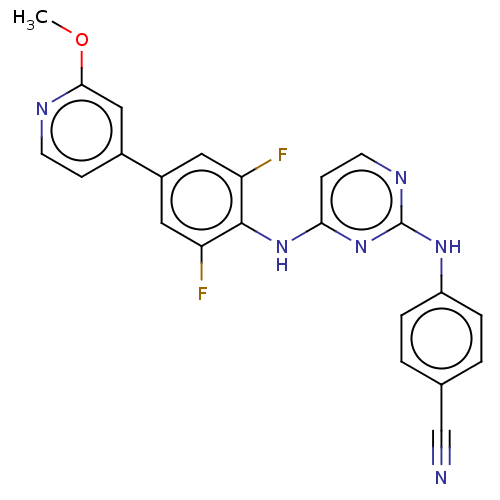
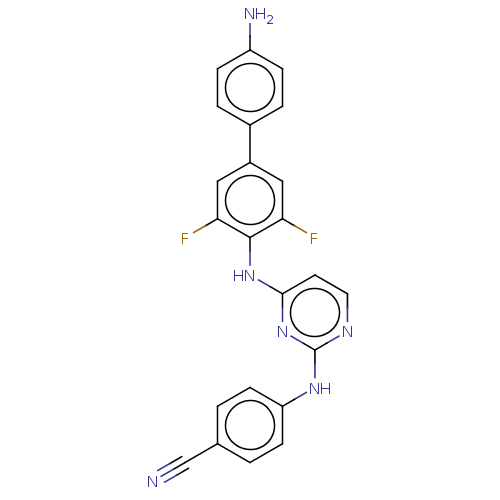
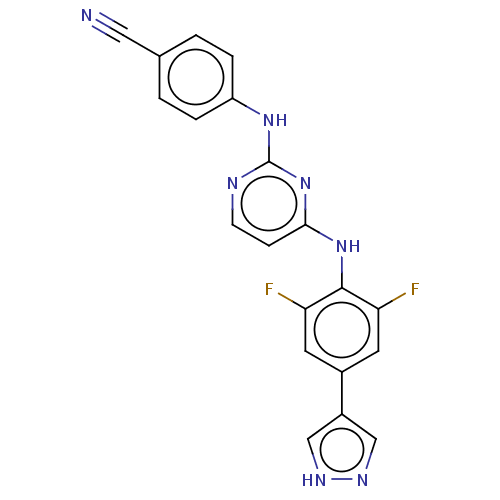
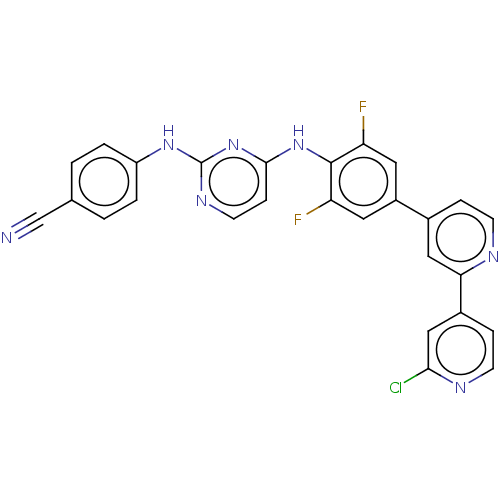
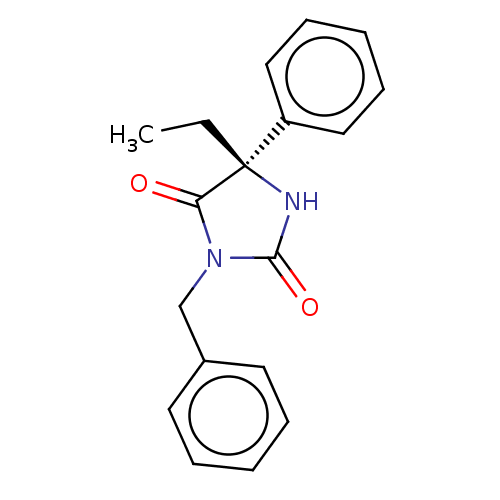
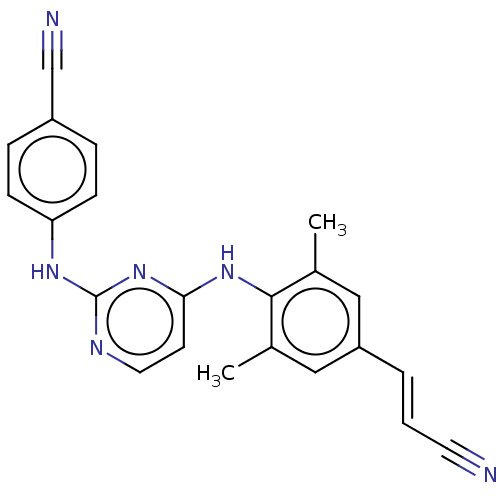
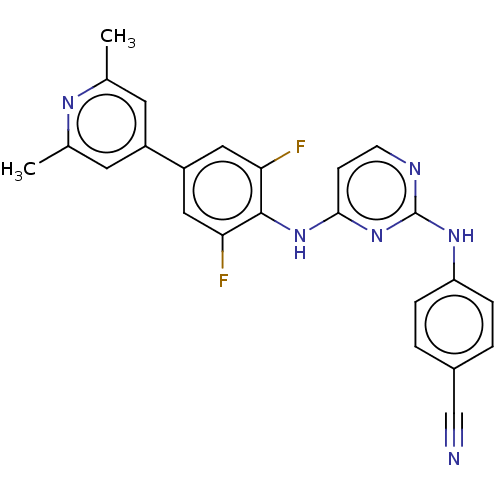
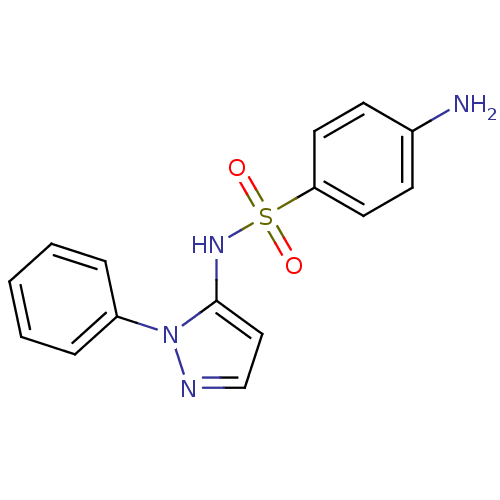
Report error Found 43 Enz. Inhib. hit(s) with all data for entry = 50014886
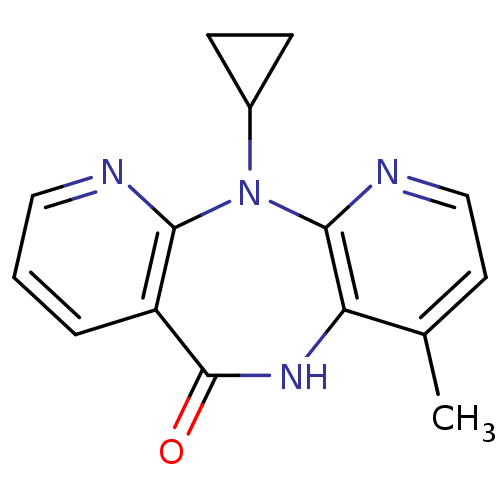
TargetReverse transcriptase/RNaseH(Human immunodeficiency virus type 1)
Fudan University
Curated by ChEMBL
Fudan University
Curated by ChEMBL
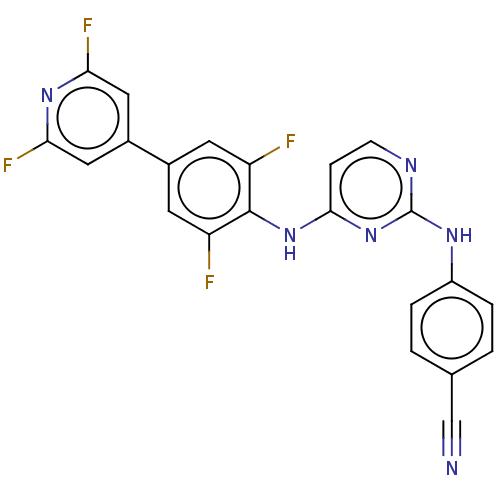
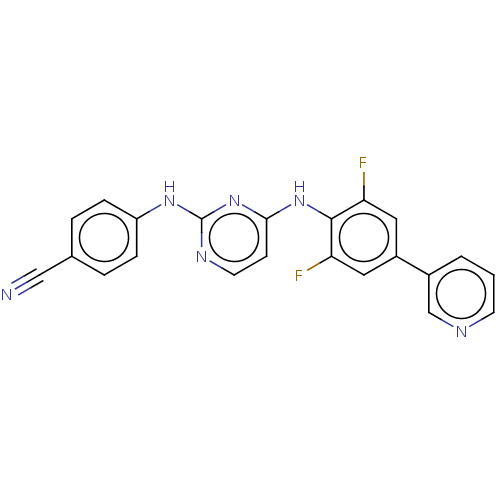
Affinity DataIC50: 5.90nMAssay Description:Inhibition of recombinant wild-type HIV1 p66/p51 reverse transcriptase assessed as inhibition of [3H]dGTP incorporation using poly (rA) as templates ...More data for this Ligand-Target Pair
TargetReverse transcriptase/RNaseH(Human immunodeficiency virus type 1)
Fudan University
Curated by ChEMBL
Fudan University
Curated by ChEMBL
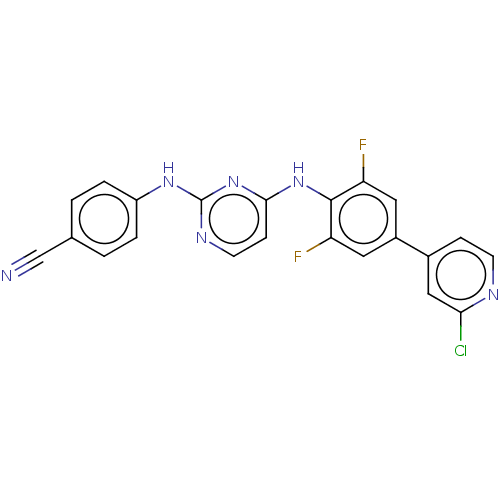
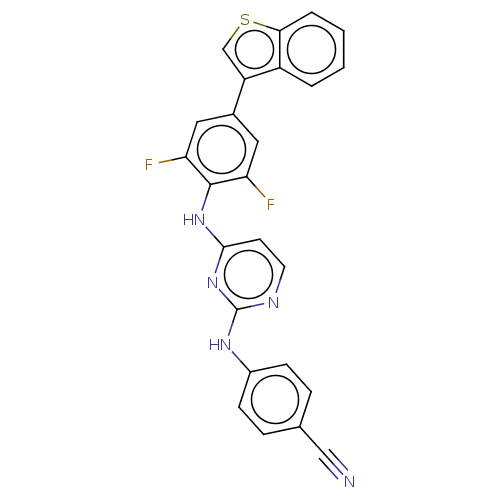
Affinity DataIC50: 7.40nMAssay Description:Inhibition of recombinant wild-type HIV1 p66/p51 reverse transcriptase assessed as inhibition of [3H]dGTP incorporation using poly (rA) as templates ...More data for this Ligand-Target Pair
TargetReverse transcriptase/RNaseH(Human immunodeficiency virus type 1)
Fudan University
Curated by ChEMBL
Fudan University
Curated by ChEMBL
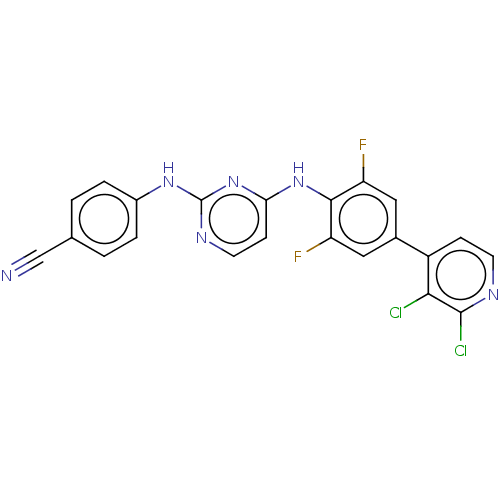
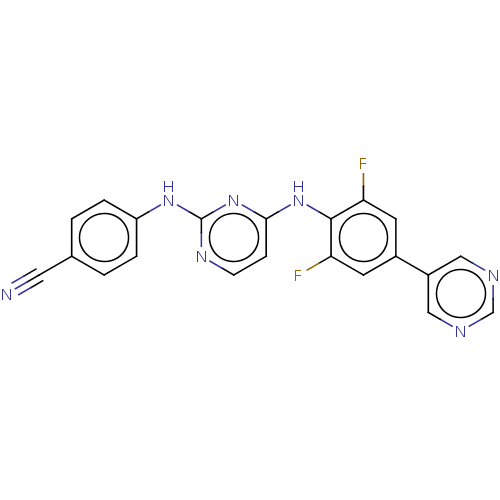
Affinity DataIC50: 8.30nMAssay Description:Inhibition of recombinant wild-type HIV1 p66/p51 reverse transcriptase assessed as inhibition of [3H]dGTP incorporation using poly (rA) as templates ...More data for this Ligand-Target Pair
TargetReverse transcriptase/RNaseH(Human immunodeficiency virus type 1)
Fudan University
Curated by ChEMBL
Fudan University
Curated by ChEMBL
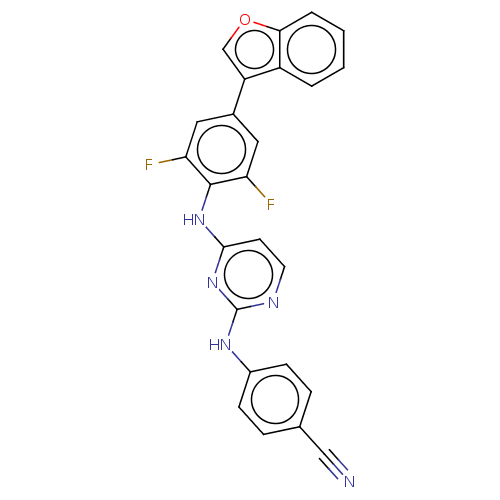
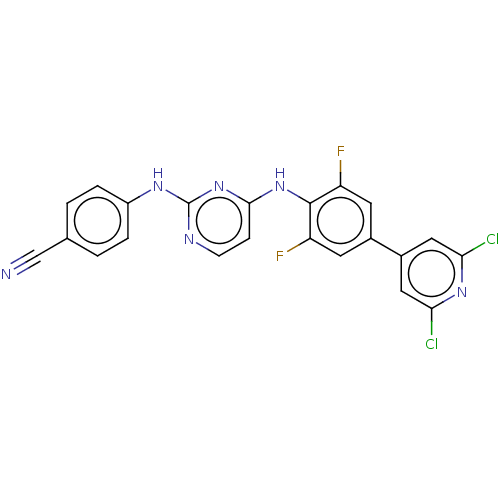
Affinity DataIC50: 9.10nMAssay Description:Inhibition of recombinant wild-type HIV1 p66/p51 reverse transcriptase assessed as inhibition of [3H]dGTP incorporation using poly (rA) as templates ...More data for this Ligand-Target Pair
TargetReverse transcriptase/RNaseH(Human immunodeficiency virus type 1)
Fudan University
Curated by ChEMBL
Fudan University
Curated by ChEMBL
Affinity DataIC50: 10nMAssay Description:Inhibition of recombinant wild-type HIV1 p66/p51 reverse transcriptase assessed as inhibition of [3H]dGTP incorporation using poly (rA) as templates ...More data for this Ligand-Target Pair
TargetReverse transcriptase/RNaseH(Human immunodeficiency virus type 1)
Fudan University
Curated by ChEMBL
Fudan University
Curated by ChEMBL
Affinity DataIC50: 10nMAssay Description:Inhibition of recombinant wild-type HIV1 p66/p51 reverse transcriptase assessed as inhibition of [3H]dGTP incorporation using poly (rA) as templates ...More data for this Ligand-Target Pair
TargetReverse transcriptase/RNaseH(Human immunodeficiency virus type 1)
Fudan University
Curated by ChEMBL
Fudan University
Curated by ChEMBL
Affinity DataIC50: 10nMAssay Description:Inhibition of recombinant wild-type HIV1 p66/p51 reverse transcriptase assessed as inhibition of [3H]dGTP incorporation using poly (rA) as templates ...More data for this Ligand-Target Pair
TargetReverse transcriptase/RNaseH(Human immunodeficiency virus type 1)
Fudan University
Curated by ChEMBL
Fudan University
Curated by ChEMBL
Affinity DataIC50: 11nMAssay Description:Inhibition of recombinant wild-type HIV1 p66/p51 reverse transcriptase assessed as inhibition of [3H]dGTP incorporation using poly (rA) as templates ...More data for this Ligand-Target Pair
TargetReverse transcriptase/RNaseH(Human immunodeficiency virus type 1)
Fudan University
Curated by ChEMBL
Fudan University
Curated by ChEMBL
Affinity DataIC50: 11nMAssay Description:Inhibition of recombinant wild-type HIV1 p66/p51 reverse transcriptase assessed as inhibition of [3H]dGTP incorporation using poly (rA) as templates ...More data for this Ligand-Target Pair
TargetReverse transcriptase/RNaseH(Human immunodeficiency virus type 1)
Fudan University
Curated by ChEMBL
Fudan University
Curated by ChEMBL
Affinity DataIC50: 11nMAssay Description:Inhibition of recombinant wild-type HIV1 p66/p51 reverse transcriptase assessed as inhibition of [3H]dGTP incorporation using poly (rA) as templates ...More data for this Ligand-Target Pair
TargetReverse transcriptase/RNaseH(Human immunodeficiency virus type 1)
Fudan University
Curated by ChEMBL
Fudan University
Curated by ChEMBL
Affinity DataIC50: 11nMAssay Description:Inhibition of recombinant wild-type HIV1 p66/p51 reverse transcriptase assessed as inhibition of [3H]dGTP incorporation using poly (rA) as templates ...More data for this Ligand-Target Pair
TargetReverse transcriptase/RNaseH(Human immunodeficiency virus type 1)
Fudan University
Curated by ChEMBL
Fudan University
Curated by ChEMBL
Affinity DataIC50: 13nMAssay Description:Inhibition of recombinant wild-type HIV1 p66/p51 reverse transcriptase assessed as inhibition of [3H]dGTP incorporation using poly (rA) as templates ...More data for this Ligand-Target Pair
TargetReverse transcriptase/RNaseH(Human immunodeficiency virus type 1)
Fudan University
Curated by ChEMBL
Fudan University
Curated by ChEMBL
Affinity DataIC50: 14nMAssay Description:Inhibition of recombinant wild-type HIV1 p66/p51 reverse transcriptase assessed as inhibition of [3H]dGTP incorporation using poly (rA) as templates ...More data for this Ligand-Target Pair
TargetReverse transcriptase/RNaseH(Human immunodeficiency virus type 1)
Fudan University
Curated by ChEMBL
Fudan University
Curated by ChEMBL
Affinity DataIC50: 16nMAssay Description:Inhibition of recombinant wild-type HIV1 p66/p51 reverse transcriptase assessed as inhibition of [3H]dGTP incorporation using poly (rA) as templates ...More data for this Ligand-Target Pair
TargetReverse transcriptase/RNaseH(Human immunodeficiency virus type 1)
Fudan University
Curated by ChEMBL
Fudan University
Curated by ChEMBL
Affinity DataIC50: 28nMAssay Description:Inhibition of recombinant wild-type HIV1 p66/p51 reverse transcriptase assessed as inhibition of [3H]dGTP incorporation using poly (rA) as templates ...More data for this Ligand-Target Pair
TargetReverse transcriptase/RNaseH(Human immunodeficiency virus type 1)
Fudan University
Curated by ChEMBL
Fudan University
Curated by ChEMBL
Affinity DataIC50: 29nMAssay Description:Inhibition of recombinant wild-type HIV1 p66/p51 reverse transcriptase assessed as inhibition of [3H]dGTP incorporation using poly (rA) as templates ...More data for this Ligand-Target Pair
TargetReverse transcriptase/RNaseH(Human immunodeficiency virus type 1)
Fudan University
Curated by ChEMBL
Fudan University
Curated by ChEMBL
Affinity DataIC50: 30nMAssay Description:Inhibition of recombinant wild-type HIV1 p66/p51 reverse transcriptase assessed as inhibition of [3H]dGTP incorporation using poly (rA) as templates ...More data for this Ligand-Target Pair
TargetReverse transcriptase/RNaseH(Human immunodeficiency virus type 1)
Fudan University
Curated by ChEMBL
Fudan University
Curated by ChEMBL
Affinity DataIC50: 37nMAssay Description:Inhibition of recombinant wild-type HIV1 p66/p51 reverse transcriptase assessed as inhibition of [3H]dGTP incorporation using poly (rA) as templates ...More data for this Ligand-Target Pair
TargetReverse transcriptase/RNaseH(Human immunodeficiency virus type 1)
Fudan University
Curated by ChEMBL
Fudan University
Curated by ChEMBL
Affinity DataIC50: 43nMAssay Description:Inhibition of recombinant wild-type HIV1 p66/p51 reverse transcriptase assessed as inhibition of [3H]dGTP incorporation using poly (rA) as templates ...More data for this Ligand-Target Pair
TargetReverse transcriptase/RNaseH(Human immunodeficiency virus type 1)
Fudan University
Curated by ChEMBL
Fudan University
Curated by ChEMBL
Affinity DataIC50: 58nMAssay Description:Inhibition of recombinant wild-type HIV1 p66/p51 reverse transcriptase assessed as inhibition of [3H]dGTP incorporation using poly (rA) as templates ...More data for this Ligand-Target Pair
TargetReverse transcriptase/RNaseH(Human immunodeficiency virus type 1)
Fudan University
Curated by ChEMBL
Fudan University
Curated by ChEMBL
Affinity DataIC50: 59nMAssay Description:Inhibition of recombinant wild-type HIV1 p66/p51 reverse transcriptase assessed as inhibition of [3H]dGTP incorporation using poly (rA) as templates ...More data for this Ligand-Target Pair
TargetReverse transcriptase/RNaseH(Human immunodeficiency virus type 1)
Fudan University
Curated by ChEMBL
Fudan University
Curated by ChEMBL
Affinity DataIC50: 87nMAssay Description:Inhibition of recombinant wild-type HIV1 p66/p51 reverse transcriptase assessed as inhibition of [3H]dGTP incorporation using poly (rA) as templates ...More data for this Ligand-Target Pair
TargetReverse transcriptase/RNaseH(Human immunodeficiency virus type 1)
Fudan University
Curated by ChEMBL
Fudan University
Curated by ChEMBL
Affinity DataIC50: 96nMAssay Description:Inhibition of recombinant wild-type HIV1 p66/p51 reverse transcriptase assessed as inhibition of [3H]dGTP incorporation using poly (rA) as templates ...More data for this Ligand-Target Pair
TargetReverse transcriptase/RNaseH(Human immunodeficiency virus type 1)
Fudan University
Curated by ChEMBL
Fudan University
Curated by ChEMBL
Affinity DataIC50: 110nMAssay Description:Inhibition of recombinant wild-type HIV1 p66/p51 reverse transcriptase assessed as inhibition of [3H]dGTP incorporation using poly (rA) as templates ...More data for this Ligand-Target Pair
Affinity DataIC50: 256nMAssay Description:Inhibition of human CYP2C19 using S-mephenytoin as substrate in presence of NADPH incubated for 15 to 45 minsMore data for this Ligand-Target Pair
Affinity DataIC50: 335nMAssay Description:Inhibition of human CYP2C19 using S-mephenytoin as substrate in presence of NADPH incubated for 15 to 45 minsMore data for this Ligand-Target Pair
TargetReverse transcriptase/RNaseH(Human immunodeficiency virus type 1)
Fudan University
Curated by ChEMBL
Fudan University
Curated by ChEMBL
Affinity DataIC50: 336nMAssay Description:Inhibition of recombinant wild-type HIV1 p66/p51 reverse transcriptase assessed as inhibition of [3H]dGTP incorporation using poly (rA) as templates ...More data for this Ligand-Target Pair
Affinity DataIC50: 346nMAssay Description:Inhibition of human CYP2C9 using tolbutamide as substrate in presence of NADPH incubated for 15 to 45 minsMore data for this Ligand-Target Pair
TargetReverse transcriptase/RNaseH(Human immunodeficiency virus type 1)
Fudan University
Curated by ChEMBL
Fudan University
Curated by ChEMBL
Affinity DataIC50: 580nMAssay Description:Inhibition of recombinant wild-type HIV1 p66/p51 reverse transcriptase assessed as inhibition of [3H]dGTP incorporation using poly (rA) as templates ...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+3nMAssay Description:Inhibition of human CYP1A2 using phenacetin as substrate in presence of NADPH incubated for 15 to 45 minsMore data for this Ligand-Target Pair
Affinity DataIC50: 2.00E+3nMAssay Description:Inhibition of human CYP3A4 using testosterone as substrate in presence of NADPH incubated for 15 to 45 minsMore data for this Ligand-Target Pair
Affinity DataIC50: 2.00E+3nMAssay Description:Inhibition of human CYP3A4 using midazolam as substrate in presence of NADPH incubated for 15 to 45 minsMore data for this Ligand-Target Pair
Affinity DataIC50: 2.17E+3nMAssay Description:Inhibition of human CYP3A4 using testosterone as substrate in presence of NADPH incubated for 15 to 45 minsMore data for this Ligand-Target Pair
Affinity DataIC50: 3.41E+3nMAssay Description:Inhibition of human CYP2D6 using dextromethorphan as substrate in presence of NADPH incubated for 15 to 45 minsMore data for this Ligand-Target Pair
Affinity DataIC50: 5.00E+3nMAssay Description:Inhibition of human CYP2D6 using dextromethorphan as substrate in presence of NADPH incubated for 15 to 45 minsMore data for this Ligand-Target Pair
Affinity DataIC50: 9.11E+3nMAssay Description:Inhibition of human CYP1A2 using phenacetin as substrate in presence of NADPH incubated for 15 to 45 minsMore data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+4nMAssay Description:Inhibition of human CYP2C9 using tolbutamide as substrate in presence of NADPH incubated for 15 to 45 minsMore data for this Ligand-Target Pair
Affinity DataIC50: 2.20E+4nMAssay Description:Inhibition of human CYP2C19 using S-mephenytoin as substrate in presence of NADPH incubated for 15 to 45 minsMore data for this Ligand-Target Pair
Affinity DataIC50: 3.00E+4nMAssay Description:Inhibition of human CYP3A4 using testosterone as substrate in presence of NADPH incubated for 15 to 45 minsMore data for this Ligand-Target Pair
Affinity DataIC50: 3.00E+4nMAssay Description:Inhibition of human CYP3A4 using midazolam as substrate in presence of NADPH incubated for 15 to 45 minsMore data for this Ligand-Target Pair
Affinity DataIC50: 3.00E+4nMAssay Description:Inhibition of human CYP1A2 using phenacetin as substrate in presence of NADPH incubated for 15 to 45 minsMore data for this Ligand-Target Pair
Affinity DataIC50: 3.00E+4nMAssay Description:Inhibition of human CYP2C9 using tolbutamide as substrate in presence of NADPH incubated for 15 to 45 minsMore data for this Ligand-Target Pair
Affinity DataIC50: 3.00E+4nMAssay Description:Inhibition of human CYP2D6 using dextromethorphan as substrate in presence of NADPH incubated for 15 to 45 minsMore data for this Ligand-Target Pair
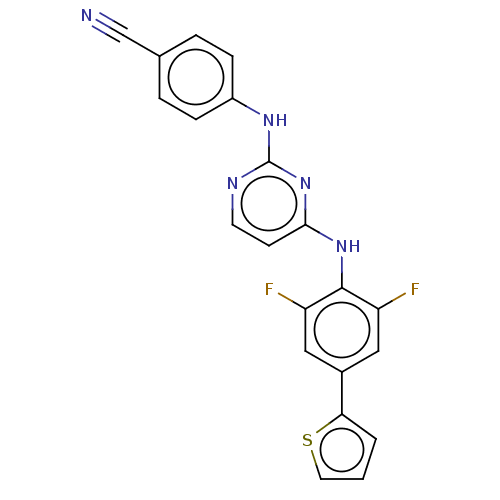
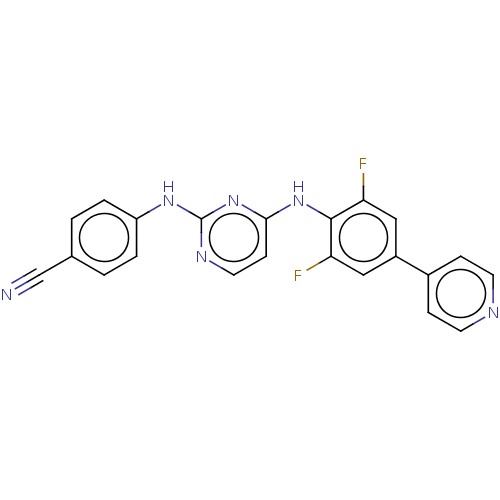
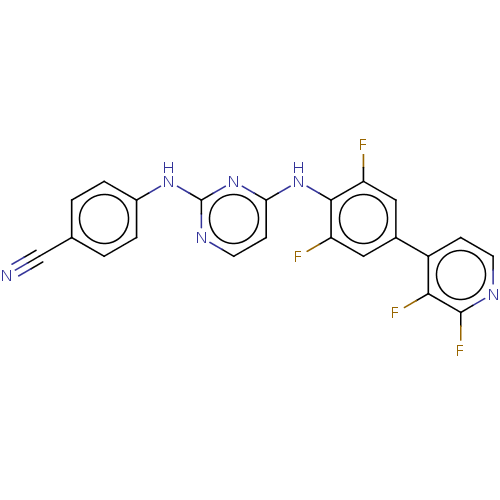
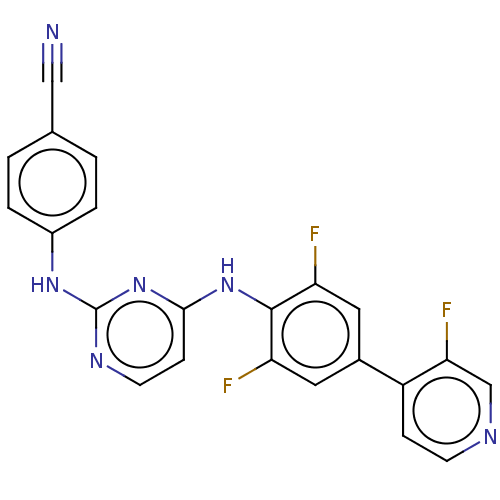
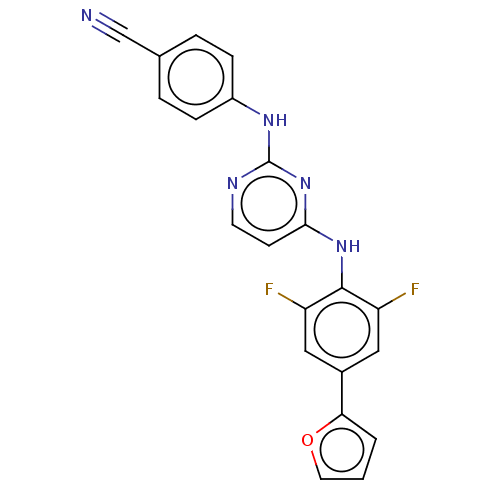
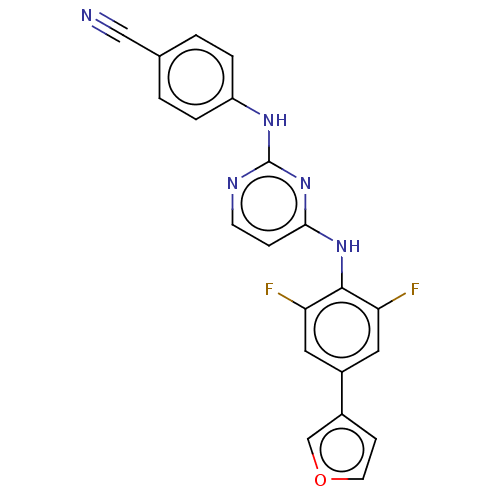
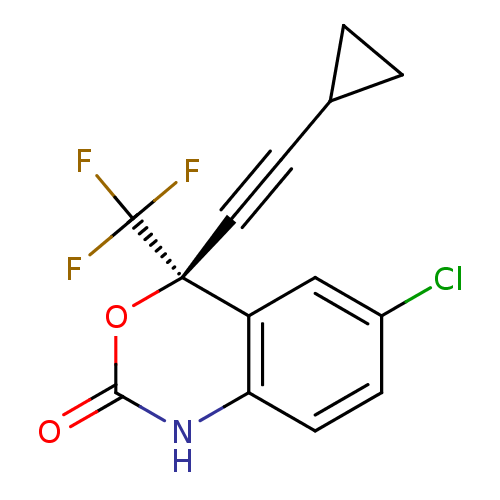
 3D Structure (crystal)
3D Structure (crystal)